New lipid class may shed light on causes of dental disease
Myriad microbial species live in the mouth, and an imbalance in their relative numbers can lead to dental disease. In chronic adult periodontitis, a destructive inflammatory disease, researchers have found Porphyromonas gingivalis, of the Bacteroidetes phylum, in increased numbers between the tooth and gum.
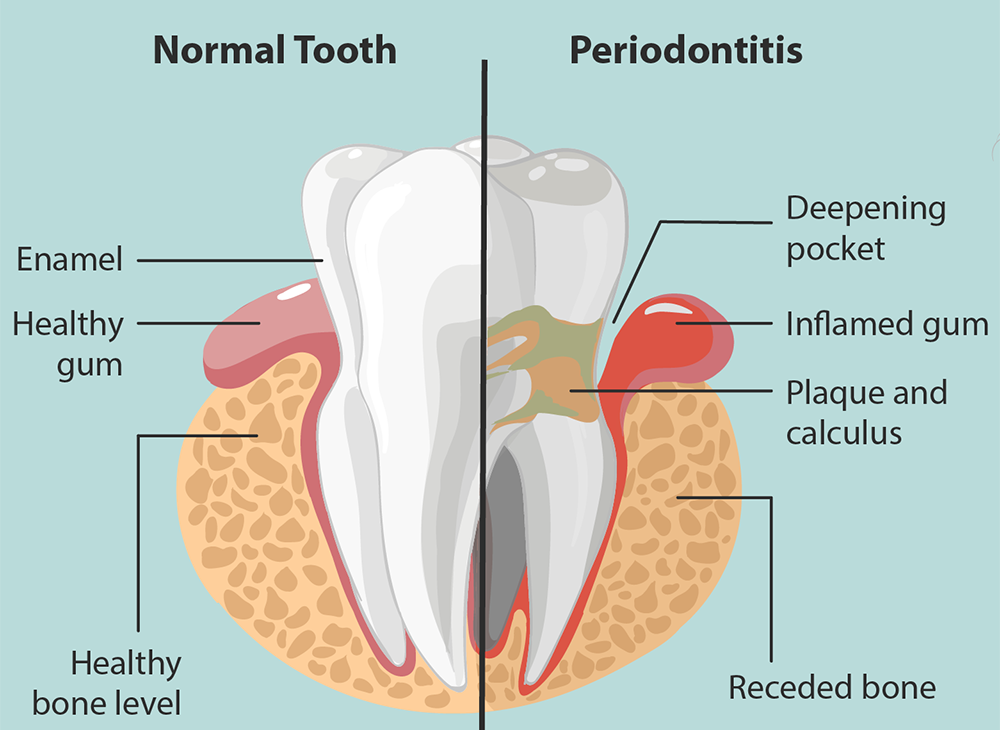
Gingivitis precedes development of periodontitis, which occurs when the inflammatory lesion of gingivitis becomes destructive, resulting in bone loss and detachment. Researchers believe virulence factors of P. gingivalis contribute to this progression. These factors include lipopolysaccharide that may promote bone resorption, expression of hairlike fimbriae that may help bacteria attach to epithelial cells between tooth and gum, and proteolytic enzymes that may promote tissue degradation. These factors could work together to promote destruction of tissue surrounding a diseased tooth.
Frank Nichols, a professor at the University of Connecticut School of Dental Medicine, described one of the possible mechanisms for delivering bacterial lipids to gingival tissues: “P. gingivalis can attach to the epithelium through its cell surface adhesion molecules, including fimbriae on the bacteria’s outer surface, and in so doing, it may deliver its cell wall lipids to epithelial cells through a process of lateral diffusion.”
In a new study in the Journal of Lipid Research, Nichols and colleagues describe isolating a new lipid class from lab-grown P. gingivalis using high-performance liquid chromatography, or HPLC, and mass spectrometry. “We noticed a small late-emerging lipid peak that eluted well after other lipid classes of P. gingivalis total lipids,” Nichols said.
The new lipid peak contained a group of related lipid species, with the most abundant having a negative ion mass of m/z 1256. The team had identified a new class of P. gingivalis lipid products — Lipid 1256. They later found that this lipid is elevated in diseased tissues and plaque samples from around teeth with periodontitis.
The Lipid 1256 class appears to be produced through a lipid synthetic pathway involving addition of amino acid or fatty acid components to the base glycine lipid structure, Lipid 342, with the final step being addition of diacylated phosphoglycerol to the serine dipeptide lipid class Lipid 654.
The researchers found that a neutralizing antibody blocked activation of human embryonic kidney cells transfected with human Toll-like receptor 2, or TLR2, demonstrating that Lipid 1256 engages TLR2. They also found that Lipid 1256 is a strong ligand for TLR2 when compared to the other serine/glycine lipid classes. Lipid 1256 stimulates release of two cytokines, IL-1β and TNF-α, from peripheral blood monocytes, both known to increase bone resorptive processes.
“Bacteria of the phylum Bacteroidetes are represented in large numbers in the gastrointestinal tract,” Nichols said, “so we wondered if Lipid 1256 is prevalent in those organisms.” They found that two common GI Bacteroidetes organisms produce Lipid 1256, suggesting they might affect chronic inflammatory disease of the GI tract by exposing the GI wall to these lipids.
Recovery of Lipid 1256 in the mouth and intestine means that both sources could contribute to diseases such as rheumatoid arthritis or atherosclerosis. Other researchers have found that these diseases may involve engagement of TLR2, which could occur if either the bacterial lipid or the bacteria enters the bloodstream. Future research evaluating the recovery of Lipid 1256 in human tissue samples may implicate lipids of the Bacteroidetes phylum in systemic diseases with a TLR2-dependent chronic inflammatory response.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

