From the journals: August 2018
We offer a selection of recent papers on a variety of topics from the Journal of Biological Chemistry, the Journal of Lipid Research and Molecular & Cellular Proteomics.
RNR gets some R and R
 Ribonucleotide reductase forms a distinctive ring-shaped structure when inhibited.Courtesy of Drennan lab/MIT Researchers at the Massachusetts Institute of Technology have discovered how an enzyme fundamental for DNA synthesis turns on and off, prompting ideas for how to develop new antibiotics and other drugs.
Ribonucleotide reductase forms a distinctive ring-shaped structure when inhibited.Courtesy of Drennan lab/MIT Researchers at the Massachusetts Institute of Technology have discovered how an enzyme fundamental for DNA synthesis turns on and off, prompting ideas for how to develop new antibiotics and other drugs.
Ribonucleotide reductases, or RNRs, are thought to be among the oldest enzymes in evolutionary history because they are responsible for turning the building blocks of RNA into the building blocks of DNA. Catherine Drennan, a professor of biology and chemistry at MIT, studies how this enzyme works in humans and bacteria.
In a new paper in the Journal of Biological Chemistry, Drennan’s team investigated why an RNR from Escherichia coli forms a donut-shaped structure when it’s inhibited. Introducing mutations that disrupted the formation of the ring caused the enzyme to stay turned on, resulting in overproduction of deoxyribonucleotides and increased mutation rates.
“We looked at the E. coli in vivo data published by Schaaper and coworkers at (the National Institute of Environmental Health Science) in light of our in vitro results and we went, ‘Wow. Not being able to turn off is a huge problem,’” Drennan said.
Human RNRs have been targets in studies of cancer drugs, but Drennan is excited about the possibility of targeting the formation of these ring structures in bacterial RNRs as a new antibiotic strategy.
“I’m very concerned about bacterial antibiotic resistance because I have a 10-year-old daughter, and so I’ve spent the last 10 years of my life being on different antibiotics,” Drennan said. “I think that this is a really interesting difference between bacterial and human (RNRs): they all form rings, but they form different kinds of rings. And it seems like there’s much more opportunity for specificity there, for designing small molecules that can target a bacterial RNR that would have no cross-reactivity with humans.”
— Sasha Mushegian
Why do kidney disease and heart failure correlate?
People with chronic kidney disease are at unusually high risk of also developing cardiovascular disease; in fact, a patient with non-dialysis kidney disease is more likely to die of heart failure than to develop end-stage kidney failure. However, traditional atherosclerosis risk factors contribute less strongly to cardiovascular disease in chronic kidney disease patients than in subjects with intact kidney function. Researchers still are trying to figure out how chronic kidney disease is linked to cardiovascular disease and how best to prevent it. In a recent study in the Journal of Lipid Research, Kathrin Untersteller and colleagues at Saarland University Medical Center in Germany and the Medical University of Graz in Austria undertook a detailed longitudinal study of patients with chronic kidney disease who were not on dialysis.
Altered kidney function is known to change the protein content of high-density lipoproteins, or HDL, which are inversely correlated with heart disease in the general population. Untersteller and colleagues hypothesized that that the level of HDL (known as “good cholesterol”) or its protein makeup in a patient’s serum at enrollment could predict the risk of cardiovascular disease in the next five years. The researchers concluded that, although some characteristics of HDL correlated weakly with future heart disease risk, no characteristic could be used independently to predict risk after controlling for other risk factors. The study underscores the complexity of untangling causality in clinical studies.
A role in the blood for a milk protein
The enzyme plasmin carries out important functions in blood, including breaking down fibrin clots and activating growth factors. Plasmin is produced by activating the precursor protein plasminogen, so uncontrolled plasminogen activation — caused, for example, by bacterial pathogens — contributes to clotting disorders and other disease states. In a study published in the Journal of Biological Chemistry, Alexander Zwirzitz and colleagues at the Medical University of Vienna found that lactoferrin, an immunomodulatory iron-binding glycoprotein from human milk, specifically inhibits plasminogen activation by directly binding plasminogens on cell surfaces. Because circulating lactoferrin increases during pregnancy, this newly discovered function may help explain why pregnancy comes with an increased risk of thromboembolism.
Cancer’s sweet tooth and distaste for fiber
Cancer cells are metabolically quirky. For energy, they rely on aerobic glycolysis, a relatively inefficient way of getting energy out of glucose, instead of shuttling glycolysis products into the mitochondria to finish breaking them down. Besides this widespread preference of most cancers, known as the Warburg effect, colorectal cancer cells have an extra metabolic quirk called the butyrate paradox. Whereas healthy cells in the colon depend on butyrate, a short-chain fatty acid made by bacteria in the digestive system, for a majority of their energy, cancerous cells are less able to proliferate when butyrate is available.
Researchers at China Pharmaceutical University in Nanjing reported on their studies of the metabolic changes in colorectal cancer cells in a recent paper in Molecular & Cellular Proteomics. The work zeroed in on the cells’ distaste for butyrate and preference for glycolysis. Qingran Li and colleagues used a metabolomics screen and found that cancer cells, after treatment with butyrate, tend to activate mitochondrial oxidation and stop using glycolysis products to generate new nucleotides and amino acids. The researchers showed that butyrate pushes this metabolic remodeling by binding to pyruvate kinase isoform M2, or PKM2, and activating it. Active PKM2 generates pyruvate, the starting point of the Krebs cycle. This research adds evidence to the existing hypothesis that turning up PKM2 may suppress tumor growth.
Series brings history of lipid science to life
 Since 2013, the Journal of Lipid Research has been running a series of thematic reviews about what organizer Alfred H. Merrill Jr. originally dubbed the “Living History of Lipids.”
Since 2013, the Journal of Lipid Research has been running a series of thematic reviews about what organizer Alfred H. Merrill Jr. originally dubbed the “Living History of Lipids.”
In his introduction to the series, Merrill described his motivation for starting the collection this way: “Much of what we know about lipids, and might be inclined to assume was easy to discover, arose from incredibly hard work, cleverly designed experiments, astonishing coincidences, and, sometimes, colossal accidents. This series of thematic reviews is intended to give glimpses into these stories. The authors will try to present the events and personalities as living histories where, when possible, readers will have a sense of stepping back in time.”
Thus far, the series has covered the lipid hypothesis of atherosclerosis, eight decades of bile acid chemistry, the discovery of essential fatty acids, what ApoE knockout and -in mice have taught us about atherogenesis, and early studies of arachidonic acid.
The latest installment, the sixth in the series, by Jean E. Vance of the University of Alberta, was published this spring. It is about the discovery, chemistry and biochemistry of two ubiquitous phosphoglycerolipids — phosphatidylserine and phosphatidylethanolamine.
PS and PE, as they’re known for short, captured Vance’s attention back when she was a postdoctoral researcher at the University of California, San Diego, working in the lab of Daniel Steinberg. (Steinberg, by the way, wrote the first installment of the “Living History” series.)
“My interest in what I felt were the rather neglected phospholipids, PS and PE, arose from some of my preliminary data suggesting that phospholipids could be compartmentalized into distinct pools in cells, perhaps due to specific inter-organelle lipid trafficking events,” Vance recalled. “(M)y research evolved into studying the biosynthesis, cell biology and functions of PS and PE in mammalian cells. Consequently, a major focus of my research was to understand the mechanism by which PS is transported from its site of synthesis in an ER domain — mitochondria-associated membranes, or MAM – to mitochondria for decarboxylation to PE.”
— Angela Hopp
Alzheimer’s protease curates neuron surfaces
The brains of people with Alzheimer’s disease contain many protein aggregates outside of cells, known as plaques. These mainly are made of the peptide amyloid-beta, which is released from the plasma membrane when the protease BACE1 cleaves its membrane-anchored precursor protein. Because amyloid-beta cannot be produced without BACE1, numerous BACE1 inhibitors have been tested or are in clinical trials as Alzheimer’s therapy.
In a recent article in Molecular & Cellular Proteomics, Julia Herber and colleagues at the German Center for Neurodegenerative Diseases described how they used a targeted surface glycoproteomics method to observe the effects of BACE1 inhibition. By labeling glycosylated membrane proteins, the researchers showed that BACE1 inhibition increases the abundance of unprocessed amyloid precursor protein but also increases other BACE1 substrates and even nonsubstrate proteins. This suggests that the inhibitor may exert unanticipated side effects by remodeling neuronal surface proteomes.
In another blow to BACE1 inhibition, major drug companies Merck and Pfizer killed BACE1 inhibitor trials this year because the drugs showed no benefit. But other companies, including Eli Lilly and Novartis, still have inhibitors in testing in the clinic.
Regulating endosomal pH
Numerous disorders, including neurodegenerative diseases, are associated with defective pH regulation in the endolysosomal pathway. Hari Prasad and Rajini Rao from the Johns Hopkins University School of Medicine performed a meta-analysis of factors affecting the regulation of an endosomal ion exchanger. They found, conserved across yeast, flies and mammals, that these exchangers (and therefore endosomal pH) were regulated transcriptionally by inhibition of a histone deacetylase, or HDAC, in response to nutrient limitation. Pharmacologically increasing expression of the HDAC corrected endosomal pH and improved clearance of amyloid proteins in a cell model of Alzheimer’s disease. Their study was published in the Journal of Biological Chemistry.
A pathogen grabs back
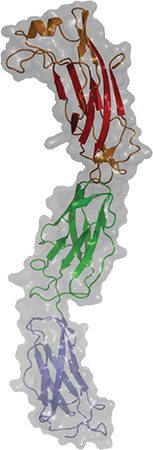 A newly published crystal structure of invasin D is part of a study of how the pathogen Yersinia pseudotuberculosis invades its host.COURTESY OF ANDREA SCRIMA Some pathogens seem impressively bold when evading the immune system, engaging with the immune system’s weapons directly to disarm them. An interdisciplinary team at the Helmholtz Centre for Infection Research published a study in the Journal of Biological Chemistry that describes how a bacterial protein grabs on to host antibodies, possibly to avoid being attacked by them.
A newly published crystal structure of invasin D is part of a study of how the pathogen Yersinia pseudotuberculosis invades its host.COURTESY OF ANDREA SCRIMA Some pathogens seem impressively bold when evading the immune system, engaging with the immune system’s weapons directly to disarm them. An interdisciplinary team at the Helmholtz Centre for Infection Research published a study in the Journal of Biological Chemistry that describes how a bacterial protein grabs on to host antibodies, possibly to avoid being attacked by them.
Andrea Scrima’s research team was interested in a group of virulence factors called invasins, which assist pathogens of the genus Yersinia (including the plague bacterium Yersinia pestis) in invading the host. In the new study, they focused on invasin D from the zoonotic pathogen Yersinia pseudotuberculosis.
“Not much was known about this protein except for its DNA sequence. Nothing was known about its particular function,” Scrima said.
The team solved the crystal structure of the protein, investigated when it’s expressed during infection and looked for its interaction partners. They found that invasin D bound the immunoglobulin A and G antibodies.
But rather than this interaction being the result of the host attacking the bacterium, it was actually the bacterial protein that was intercepting the antibody — a bit like grabbing someone by the wrist to avoid being grabbed by their hand.
The study required collaboration among groups with diverse expertise, which Scrima credits for the insight into this unusual form of immune evasion.
“This, in my opinion, is a very good example of a great and interesting interdisciplinary collaboration that shows that the whole is more than the sum of its parts,” Scrima said.
An enzyme’s strange residues
Lysosomal phospholipase A2, or LPA2, is an unusual enzyme. In addition to interacting with a wide range of substrates, LPA2 can act as either an acyltransferase or a phospholipase. It also plays a role in host defense mechanisms, and, in contrast to many neutrally active phospholipases, is most active in highly acidic pH.
To figure out why LPA2 has such strange properties, researchers at the University of Michigan honed in on Asp13, a recently identified acidic residue on the enzyme. Their paper in the Journal of Lipid Research describes how, when they substituted Asp13 for a variety of amino acids, the researchers found that LPA2 became most active at neutral pH and was less able to interact with the mono- and double-unsaturated acyl chains that it previously had favored.
Six degrees of dopamine receptor mutation
Dopamine disturbances are thought to play a role in multiple psychiatric disorders. Freja Herborg and colleagues at the University of Copenhagen systematically compared the effects of six rare mutations, identified in patients with diverse neuropsychiatric conditions, on the function of the dopamine active transporter, or DAT. They found that all six of the variants had unique structural and functional changes but that disruptions in ion conductance were a common theme that might underlie disruption of dopamine transport in neuropsychiatric disease. The study was published in the Journal of Biological Chemistry.
Pore forming, proteome remodeling
It’s a tale nearly as old as genetic information: One set of cells would like to continue its daily business of protein synthesis and replication, while another would like to sabotage those mechanisms for its own gain. When the pathogen Listeria monocytogenes, of foodborne infamy, finagles its way inside epithelial cells in the human intestines, the bacterium deploys the pore-forming toxin Listeriolysin O, or LLO, which interferes with the proteins synthesized by the infected cell. This ultimately results in cell death by creating holes in the cell membranes.
In a paper in Molecular & Cellular Proteomics, researchers at the Pasteur Institute in Paris describe a proteomics analysis of human epithelial cells treated with LLO, in which they found that the toxin acts exclusively by altering host proteins through post-translational modifications involving ubiquitin rather than affecting transcriptional activity of underlying genes. They also found that a similar toxin, Perfringolysin O, acts through proteome remodeling.
G protein gadgets
G protein–coupled receptors mediate many important signaling processes, but their activity is difficult to study in live cells. Nevin Lambert’s lab at the University of Georgia developed miniaturized G proteins fused to reporter probes as tools to monitor GPCR activation directly, precisely and sensitively. These constructs can be adapted to diverse assay formats and be used with standard laboratory equipment, making them useful for interrogating many signaling pathways and drug targets. The research was published in the Journal of Biological Chemistry.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.


.jpg?lang=en-US&width=300&height=300&ext=.jpg)

