A mechanism for remdesivir activity and a platform to test other antivirals
Researchers who showed in February that remdesivir blocks coronavirus polymerases are reporting in the Journal of Biological Chemistry that the finding also applies to the novel coronavirus —and they think they can explain the molecular mechanism for how the drug gums up the virus’ gears.

In their earlier paper, graduate student Calvin Gordon and professor Matthias Götte of the University of Alberta and co-authors demonstrated that remdesivir can inhibit the viral replicase of the coronavirus that causes Middle East Respiratory Syndrome, or MERS. The study included co-authors from Gilead Sciences, the company that makes the drug.
In the ensuing months, remdesivir, an adenosine-mimicking nucleotide analog developed to treat other viruses, has emerged as the most promising small-molecule treatment for COVID-19. In late April, the National Institute for Allergy and Infectious Diseases reported qualified success of a clinical trial of the drug, and the FDA rapidly issued emergency use authorization for the drug.
Now, using enzymes from the novel coronavirus SARS-nCoV-2 itself instead of the related virus that causes MERS, the research team has shown that remdesivir blocks the SARS-nCoV-2 replicase in a test tube. They also examined a selection of other nucleotide analogs to understand more about whether and how each might work against infection.
The Götte lab studies viral enzymology, and Götte has developed a system for expressing large quantities of viral polymerases from insect cells to enable rapid enzymology studies, informed by a World Health Organization pandemic-preparedness blueprint, which regularly updates a list of the most concerning pathogens. “Coronaviruses like SARS and MERS were always on the list,” Götte said.
Most graduate students don’t finish a paper in their first year in the lab — let alone two papers. Calvin Gordon, a first-year graduate student at the University of Alberta in Edmonton, joined the lab in September 2019. When he began the MERS study in October, the risk of a pandemic was still theoretical. By the time the authors were ready to write up their finding that the MERS polymerase easily incorporates remdesivir but stumbles to a halt a short time later, the first cases of COVID-19 had been reported in Hubei province, China. And by the time they published the discovery, Italy was in lockdown and cases had been reported in both the U.S. and Canada.
As soon as the first SARS-CoV-2 genome was published in January, Egor Tchesnokov, a research associate in the lab and co–first author on the new paper, got to work cloning it into their expression system and preparing to test remdesivir’s effect on those enzymes. Their new study, published in April, confirms that, as it did in MERS, the drug works against the enzyme complex the novel coronavirus uses to replicate its genome. This time, thanks to a molecular modeling study run by Gilead’s Jason Perry, the authors are able to explain how it happens.
Using the crystal structure of the polymerase enzyme from another coronavirus — as it happened, researchers from China and Australia published a cryo-EM structure of the novel coronavirus’ replicase on the same day as the second Götte study — and a crystal structure showing how a hepatitis virus enzyme interacts with remdesivir, Perry and other researchers at Gilead put together a computational model of how remdesivir fits into the active site of the novel coronavirus’ replicase.
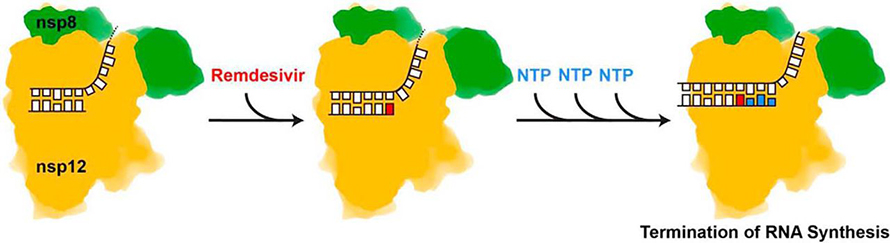
“As the enzyme incorporates one, two, three more nucleotides, the incorporated remdesivir moves back, so to speak,” Götte said. As it reaches the third position away from the enzyme’s active site, the drug encounters — or causes — a blockage. Because of its unusual 1' modification, it crashes into a specific amino acid, a serine conserved among most coronavirus polymerases, which prevents the enzyme from moving one step forward to incorporate the next RNA base.
“It’s not perfect chain termination; there’s a little read-through,” Götte said. “The specific wish would be to see whether there’s any other nucleotide (analog) out there that would cause a stronger termination.”

To find out, the researchers assessed the effect of a variety of other known nucleotide analogs — some of which are in clinical trials — on the viral polymerase. This allowed them both to pinpoint the effect of the drug on a chemical modification at the 1' in its ribose by comparing the features of molecules that did and did not inhibit the polymerase and to make some predictions about other nucleotide analog drug candidates.
For example, they found that, unlike polymerases from other viruses, the enzyme did not recognize drugs with a 2' ribose modification as RNA building blocks. For a drug to inhibit the viral replicase, the replicase must recognize it as a building block of new RNA, and for realistic dosing, it’s best if the enzyme picks up the drug more easily than the drug’s natural counterpart. The researchers found that the SARS-CoV-2 polymerase does not prefer a Gilead drug for hepatitis, sofosbuvir, to its natural counterpart, UTP. In a competition assay, Götte said, “UTP wins, by far. So it is very unlikely based on our data that we would expect potent antiviral activity from sofosbuvir.”
They also investigated the influenza drug favipiravir, which is being tested in fifteen clinical trials around the world for efficacy as a COVID-19 treatment. Like sofosbuvir and remdesivir, favipiravir is a nucleotide analog that has been shown to inhibit other RNA viruses. However, as with sofosbuvir, the SARS-CoV-2 polymerase showed strong selectivity for the natural ribonucleotide over favipiravir, raising some concern about whether favipiravir will have antiviral activity against the new coronavirus. Götte said, “I think it’s very important to reconcile the biochemical data with antiviral data.”
Whether or not remdesivir proves to be an effective treatment for COVID-19, the researchers hope their enzyme platform will help speed up assessment of novel drug candidates as they are developed. They also plan to seek therapeutic combinations of nucleotide inhibitors and drugs with other mechanisms, which could reduce the chances that resistance will develop, Götte said.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

