How an oral antiviral drug confuses the replication machinery of SARS-CoV-2
A University of Alberta virology lab has uncovered how an oral antiviral drug works to attack the SARS-CoV-2 virus. The findings were published recently in the Journal of Biological Chemistry.
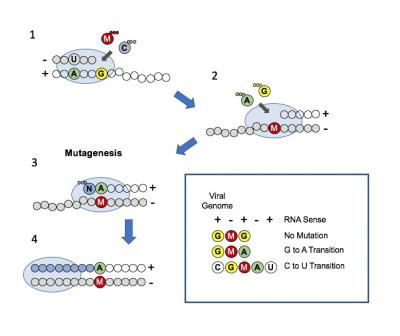
(oval)–mediated nucleotide incorporation into the RNA primer
(grey circles)/template (white circles). Plus and minus signs indicate
RNA sense. A, C, G and U refer to natural nucleotide bases, and M
refers to the active forms of molnupiravir. Three small circles indicate
the triphosphate form of nucleotides. (1) Molnupiravir competes with CTP
for incorporation during synthesis of the negative RNA strand
(copy of the genome). (2) When embedded in the template, molnupiravir
base-pairs with either ATP or GTP. (3) Incorporation of ATP results
in mutagenesis. (4) The mutation is fixed as RNA synthesis continues.
The boxed information provides a summary of events that lead to drug-induced
G to A and C to U transition mutations.
The researchers demonstrated the underlying mechanism of action by which the antiviral drug molnupiravir changes the viral genome, a process known as excessive mutagenesis or error catastrophe.
Matthias Götte is a professor and chair of the medical microbiology and immunology department in the university’s Faculty of Medicine & Dentistry and a member of the Li Ka Shing Institute of Virology. “The polymerase, or replication engine of the virus, mistakes molnupiravir molecules for the natural building blocks required for viral genome replication and mixes them in,” Gotte said. “It causes the polymerase to make sloppy copies — nonsense genomes that are useless and not viable.”
In clinical trials for efficacy, molnupiravir eliminated SARS-CoV-2 infectivity in newly diagnosed patients after five days of treatment. The drug is taken as a pill, making it much easier to administer than other approved treatments such as remdesivir or monoclonal antibodies, which must be given intravenously.”
“Our work to demonstrate that the effect of the drug is indeed mediated by the viral polymerase is reassuring, because if the drug somehow generates mistakes in the virus and you don’t know how it happens, there could be other mechanisms at work that could also harm the cell. Still, the safety of the drug for COVID-19 patients remains to be evaluated and monitored.”
The ongoing hunt for a weapon against pandemics
The active form of molnupiravir first was identified as a broad-spectrum antiviral at Emory University in Atlanta, Georgia. In 2003, it was developed as a treatment for chronic hepatitis C, but it was dropped due to possible side effects associated with long-term use. The drug then was developed as an influenza antiviral, because the course of treatment for flu is much shorter. The focus of testing switched to SARS-CoV-2 after the COVID-19 pandemic emerged. The drug now is being developed in partnership by Merck and Ridgeback Biotherapeutics.
Merck has made deals with five generic drugmakers in India to make molnupiravir, and at least one of them has applied for approval to use it on an emergency basis, as at least 350,000 new infections are diagnosed in that country every day and vaccination levels are low.
Götte and his team previously uncovered the mechanisms of action for remdesivir, a now-approved treatment that inhibits replication of the SARS-CoV-2 virus, and baloxavir, an influenza drug.
Next, they will test molnupiravir’s mechanism of action against the polymerases of some of the other viruses the World Health Organization has identified as having high epidemic potential.
“All are recognized as emerging pathogens where we need to develop countermeasures,” Götte said. “We need to be prepared with broad-spectrum antivirals that can serve as a first line of defense.
“Even once vaccines are developed, we can’t get them into all the arms at once,” he said. “To really fight outbreaks and epidemics, one tool is unlikely to be sufficient.”
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

