Arginine tango
As a means to evade the host immune response, S. aureus uses an enzyme called oleate hydratase, or OhyA, to inactivate antimicrobial unsaturated fatty acids in the membrane that would otherwise inhibit bacterial growth. Research scientists at St. Jude Children’s Research Hospital reported today the structure and catalytic mechanism of OhyA.
Christopher Radka of St. Jude’s Department of Infectious Diseases describes the research during a presentation Tuesday at 2 p.m. EDT at the 2021 ASBMB Annual Meeting, held in conjunction with the Experimental Biology conference.
Radka and colleagues used X-ray crystallography to determine the structure of OhyA. Solving and evaluating multiple OhyA crystal structures highlighted a coordinated dance that occurs between key arginine residues and the unsaturated fatty acid substrate in the active site of the enzyme, a process facilitated by the nucleotide cofactor FAD.
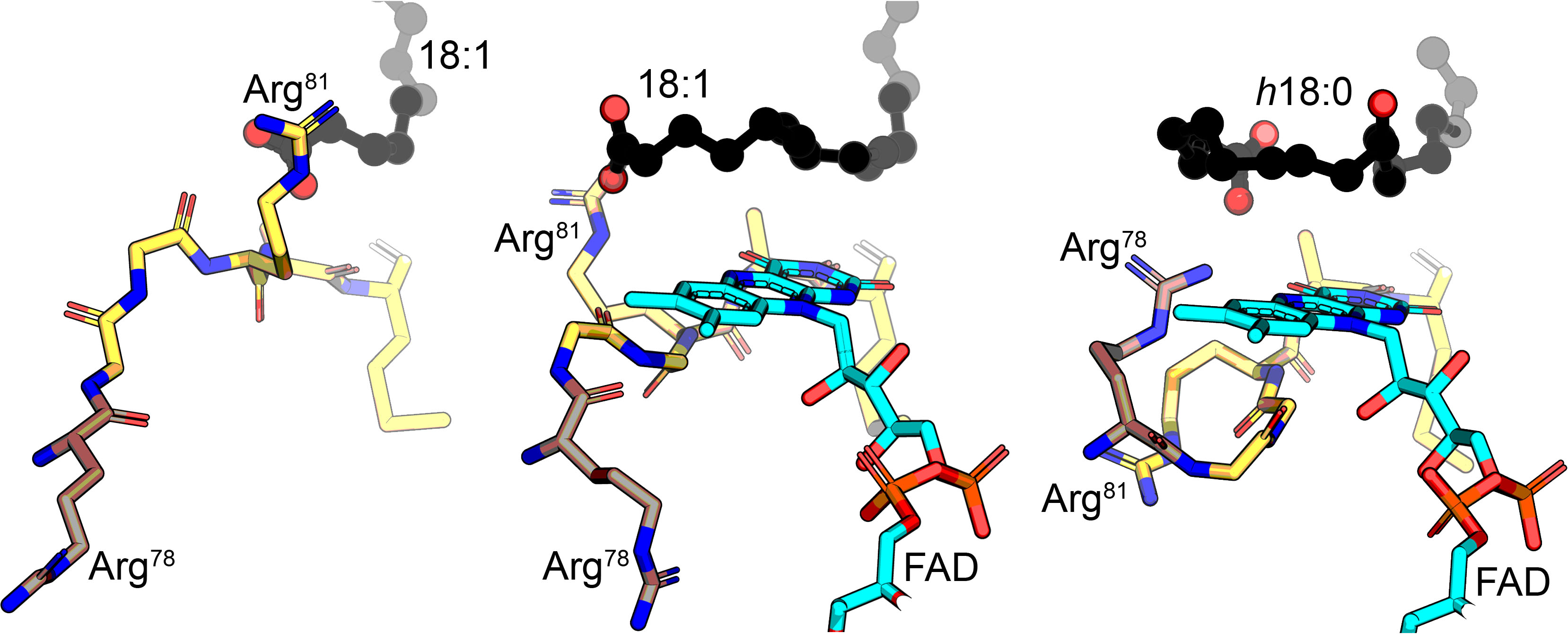
In this dance, the substrate is first guided into the binding tunnel by the oleate carbonyl of OhyA, then encounters its first arginine dance partner (Arg81) at the entrance of the active site. FAD binding then triggers the rotation of Arg81 that guides the fatty acid as it curls into the active site. After catalysis, a second arginine (Arg78) rotates behind the fatty acid carboxyl to release the hydroxylated product from the active site.
“What’s novel about the (active site) is how these conserved arginines guide the substrate through the donut-shaped active site,” Radka said. “Here, the arginines dance like two partners in a tango.”
This highly choreographed dance controls how the fatty acid substrate moves into and out of the active site. “In this coordinated tango at the active site, the FAD is the dramatic third character whose role is to come in and advance the dance so the chemistry can occur,” Radka said.
In this reaction, FAD remains oxidized and unconsumed. This quality is advantageous for industrial biotechnology research looking to use OhyA; FAD-dependent reactions often consume FADH2 and require continued starting product, which can be costly.
Future goals for this research include determining the structural elements required for S. aureus OhyA to remove antimicrobial fatty acids from the membrane.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

