A deeper insight into phospholipid biosynthesis in Gram-positive bacteria
Gram-positive bacteria can cause serious and sometimes fatal inflammatory diseases including skin infections, pharyngitis, pneumonia, endocarditis, myocarditis, meningitis, rheumatic fever and septicemia. These bacteria synthesize phosphatidic acid, the central precursor of membrane phospholipids, using an unusual acyl-phosphate intermediate in a three-step pathway mediated by three phospholipid synthesis enzymes: PlsX, PlsY and PlsC.
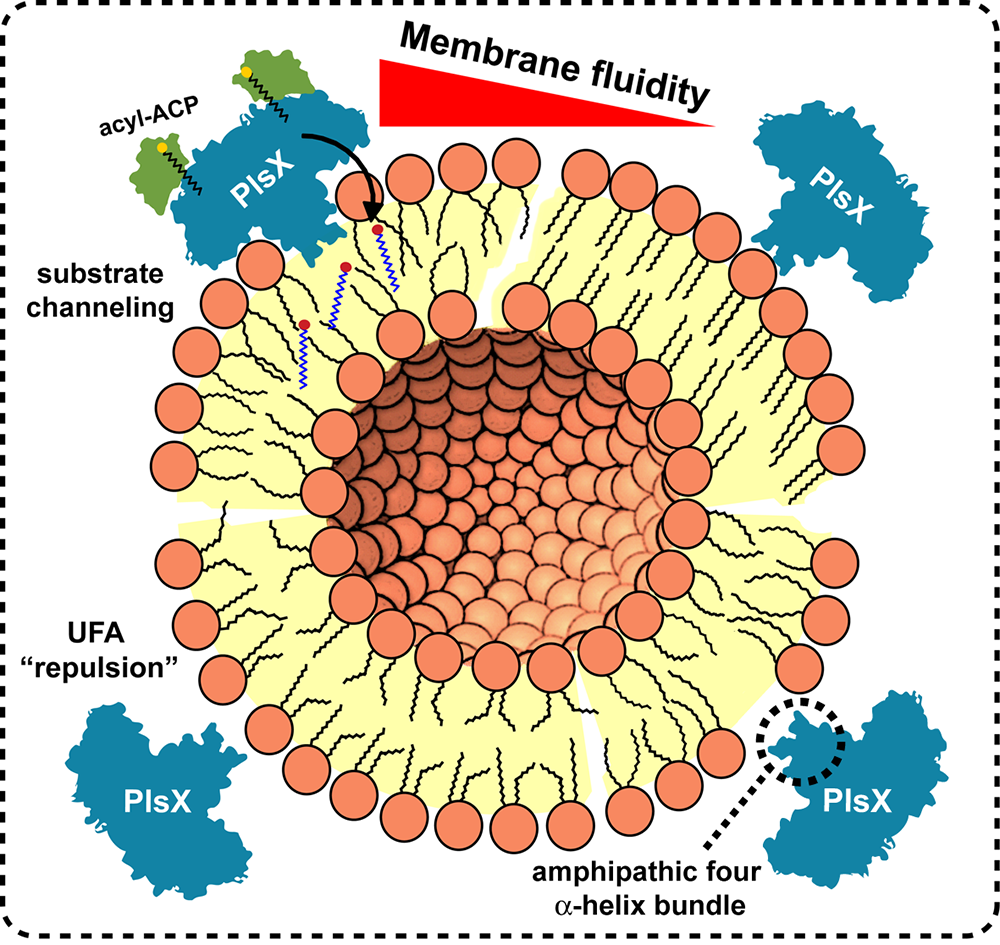
PlsX is a peripheral membrane transacylase that catalyzes the conversion of acyl-acyl carrier protein to acyl-phosphate and helps coordinate fatty acid and phospholipid biosynthesis. PlsX binds and inserts directly to lipid bilayers, a process mediated by an amphipathic four alpha-helical bundle subdomain that protrudes from the main core of the enzyme.
In disentangling this binding and insertion of PlsX, our lab has found that PlsX membrane binding is mediated by phospholipid charge, whereas unsaturation of fatty acids and membrane fluidity influence membrane insertion. Superficial access to the membrane is not sufficient to ensure efficient delivery of the acyl-phosphate from PlsX to the acyltransferase partner PlsY, which depends on proper and stable insertion of PlsX in the membrane. Such substrate channeling can make this metabolic pathway more efficient and prevents the release of unstable intermediates, protecting them from decomposition and/or diffusion through the aqueous cytoplasm.
In a recent paper in the Journal of Biological Chemistry, we propose a model in which membrane fluidity governs the membrane insertion of PlsX, which is required for the proper acyl-phosphate delivery to PlsY.
Our model breaks a paradigm in membrane biology, because the membrane fluidity and unsaturated fatty acids, or UFAs, do not seem to be holding hands. At higher membrane fluidity, PlsX inserts deeper into the membrane, but increased UFA content reduces PlsX binding and insertion.
How can PlsX sense the UFAs, and why do UFAs repel PlsX binding and insertion? We suspect the repulsion effect might be part of a mechanism to downregulate the total phospholipid synthesis observed at low growth temperature.
In membrane insertion, peripheral proteins can cluster as transient oligomers when interacting with lipid bilayers. This crowding would be intensified with increased protein radius and depth of penetration into the hydrophobic region of the membrane. PlsX mostly is distributed homogenously at the bacterial cell membrane, but PlsX foci also are transiently and randomly observed, which could represent physiological oligomeric states related to the regulation of PlsX activity.
We propose that PlsX inserts deeper into more fluid membranes (containing little or no unsaturated fatty acid content), causing oligomers, including other partners, in the form of foci at the membrane. Thus, modulating membrane insertion could regulate the substrate channeling as well as the protein crowding, regulating the phospholipid biosynthesis.
These results highlight the relevance of spatial organization for metabolic pathway functioning and tell us more about how membrane composition and protein mobility can modulate this biosynthesis route. PlsX and PlsY are present in pathogenic bacteria and absent in eukaryotes, so the results offer exciting possibilities for inhibitor design to fight antibiotic-resistant bacteria.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.


