From the journals: MCP
Protein profiles offer new insights into Duchenne muscular dystrophy. A novel fungal protease improves proteomic coverage. Expression of HLA-II shows promise for immunotherapy. Read about these recent articles in Molecular & Cellular Proteomics.
New insights into Duchenne muscular dystrophy
Duchenne muscular dystrophy, or DMD, is characterized by progressive muscle degeneration and weakness due to the alterations of the dystrophin protein that keeps muscles intact. Development of effective treatments for DMD has been challenging because researchers do not understand completely the pathological consequences resulting from loss of dystrophin.
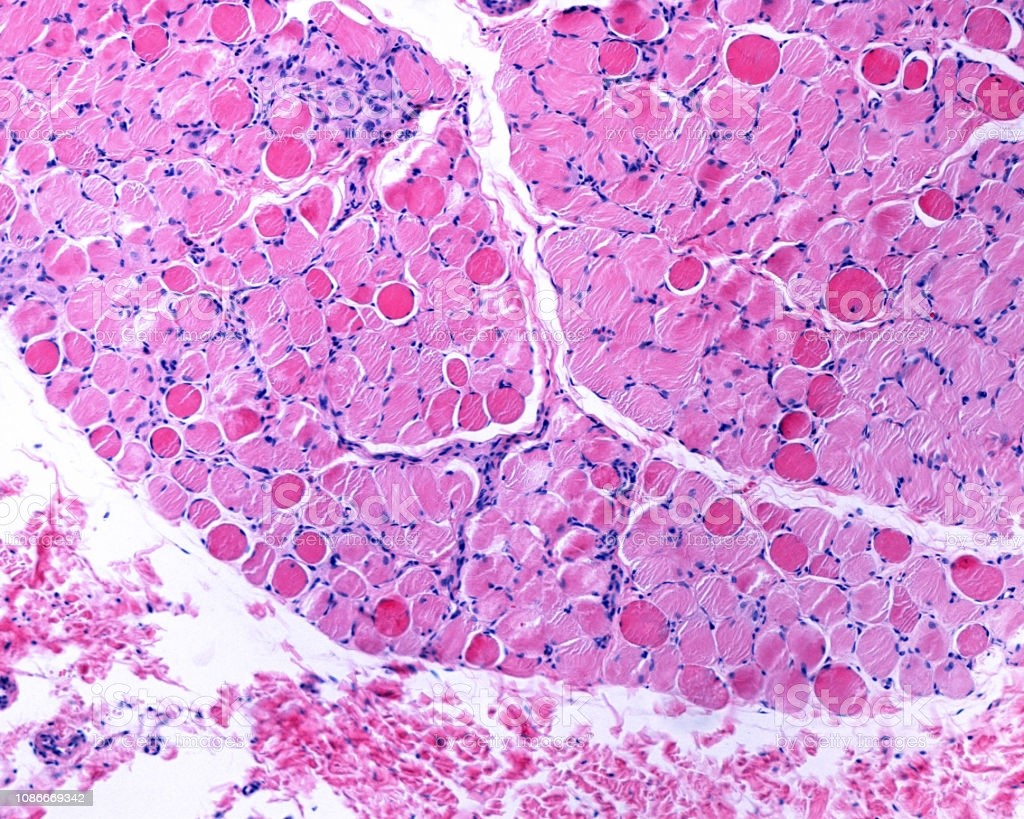
In a recent paper in the journal Molecular & Cellular Proteomics, Tirsa L.E. van Westering and researchers at the University of Oxford used mass spectrometry to identify differential protein expression in muscles at three different ages (representing the different stages of DMD pathology) from two strains of mice with genetically altered DMD and one strain of unaltered mice (control).
The protein profiles of the two types of DMD mice were found to be similar; however, numerous proteins were differentially expressed between DMD mice and unaltered mice at each of three stages of DMD pathology. These included proteins associated with the extracellular matrix and muscle function, many of which had not been reported in previous studies. Further analysis showed that the dystrophic condition is characterized by multiple perturbed pathways and crosstalk among inflammatory, metabolic and muscle growth pathways.
The researchers created a publicly available data set that reports the quantification of 4,974 proteins across the DMD mice and control mice at various disease time points — the highest number of quantified proteins in dystrophic muscle described to date. It will offer many researchers a wider perspective into the proteomic signatures characteristic of DMD and potential therapeutic targets and also will be useful for researchers studying the effects of aging on the muscle proteome.
Novel fungal protease improves proteomic coverage
Enzymatic digestion of protein into shorter peptides by sequence-specific proteases is a key step in mass spectrometry-based shotgun proteomics experiments. Trypsin has been used predominantly due to its high specificity (it cleaves exclusively after the amino acids arginine and lysine), widespread availability and ease of use.
Diana Samodova and a team at the University of Copenhagen characterized another protease, ProAlanase, isolated from the fungus Aspergillus niger, that cleaves primarily on the C-terminal side of proline and alanine amino acid residues of proteins. Compared to trypsin-only digestion, a combination of ProAlanase and tryptic digestions showed improved proteome coverage. The researchers also demonstrated ProAlanase’s broad utility in other proteomics applications in a recent paper published in the journal Molecular & Cellular Proteomics.
This study highlights the importance of investigating alternative proteases for proteomic experiments. Alternatives to trypsin will be beneficial in cases where digestion by trypsin is not ideal, such as when the content of arginine and lysine in the protein is low. Moreover, as proteases with different cleavage sites will generate different sets of peptides for the same proteome, combination digestion will yield improved results for various applications, such as identification of proteins, protein–protein interactions and post-translational modifications in various biological systems.
Expression of HLA-II shows promise for immunotherapy
When the immune system is suppressed, glioblastoma, or GBM, develops into an aggressive cancer in the brain or spinal cord, with poor prognosis. Understanding the interaction between the T-cell receptor on CD4+ T cells and HLA-II cancer-specific epitopes on GBM cells is vital for the development of immunotherapies. However, HLA-II complexes almost never are expressed naturally in GMB cells, and scientists believe their absence contributes to the poor anti-tumor immune response commonly seen in GBM.
In a recent paper in the journal Molecular & Cellular Proteomics, Greta Forlani and a team of Swiss researchers write that they discovered how to induce HLA-II expression by inserting the gene histocompatibility complex transactivator, or CIITA, into various GMB cell lines. They then used a number of techniques to uncover more than 30,000 unique HLA-II peptides in those cell lines.
If HLA-II expression can be induced successfully, it has promising therapeutic implications for HLA-II–negative tumors such as GBM. For example, mice with tumor cells injected with CIITA (which causes increased HLA-II expression) have been shown to produce robust anti-cancer immune responses. Furthermore, CIITA-expressing tumor cells will be useful for discovering antigens that can be used to develop immunotherapies.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

