From the journals: December 2019
We offer a selection of recent papers on a variety of topics from the Journal of Biological Chemistry, the Journal of Lipid Research and Molecular & Cellular Proteomics.
Chasing tails: improving structural analysis of histones
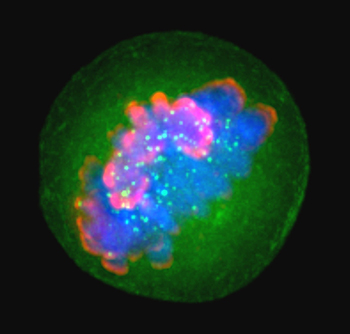 Histones such as H3, shown here in pink during the metaphase stage of mitosis, package and order DNA in structural units called nucleosomes.LAURA TRINKLE-MULCAHYHistones are essential for chromatin organization, but structural details for their N-terminal tails — which play a crucial role in epigenetic regulation by way of post-translational modifications — have proven elusive for crystallographic characterization. To obtain these structural details, researchers at the Broad Institute and the National Institute of Standards and Technology turned to hydrogen/deuterium-exchange mass spectrometry.
Histones such as H3, shown here in pink during the metaphase stage of mitosis, package and order DNA in structural units called nucleosomes.LAURA TRINKLE-MULCAHYHistones are essential for chromatin organization, but structural details for their N-terminal tails — which play a crucial role in epigenetic regulation by way of post-translational modifications — have proven elusive for crystallographic characterization. To obtain these structural details, researchers at the Broad Institute and the National Institute of Standards and Technology turned to hydrogen/deuterium-exchange mass spectrometry.
The technique, abbreviated as HX-MS, takes advantage of the tendency of hydrogens in certain proteins to trade places constantly with nearby hydrogen atoms when in aqueous solutions. When these proteins, such as histones, are exposed to heavy water that contains deuterium, a hydrogen isotope twice the mass of normal hydrogen, they wind up incorporating it in exposed positions on their surface. Once there, the deuterium atoms then can be used to probe protein conformation.
Malvina Papanastasiou and colleagues treated the four core histones — H2A, H2B, H3 and H4 — with the protease cathepsin-L, which helps improve coverage of histone tails, before subjecting them to HX-MS.
They found that cathepsin-L demonstrated unique cleavage patterns for each of the histones that were pH-dependent and that it generated overlapping N-terminal peptides about 20 amino acids long for the core histones H2A, H3 and H4. Taken together, the authors believe these results prove the protease’s suitability for the analysis of histone tail dynamics.
“Overall, this novel strategy opens new avenues for investigating the dynamic properties of histones that are not apparent from the crystal structures, providing insights into the structural basis of the histone code,” they write in their recent paper in the journal Molecular & Cellular Proteomics.
— John Arnst
Probing progesterone-producing cells in pregnancy
Extravillous trophoblasts, or EVTs, support formation of the placenta, and EVT dysfunction has been associated with pregnancy complications. However, the unique characteristics of EVTs compared to other cell types such as villous cytotrophoblasts, or vCTBs, are not well understood. Research published in the Journal of Lipid Research provides new information on the metabolic characteristics of these key cells.
Austria- and California-based researchers led by Sigrid Vondra genomically profiled EVTs and vCTBs from first-trimester human placentas and found higher cholesterol levels in EVTs as well as significant differences in genes associated with cholesterol metabolism. They found that EVTs have higher levels of the enzyme HSD3B1, which is responsible for a final step in producing progesterone (a key pregnancy hormone) from cholesterol. Through additional analyses, the researchers discovered that EVTs produced and secreted high levels of progesterone to support placental health and that they relied on their higher cholesterol levels to support this biological role. Perhaps most significantly, they found that HSD3B1 levels were decreased in placental samples from miscarriage, linking the EVTs’ progesterone production directly to pregnancy health. Further studies could provide warning signs and drug targets for healthy pregnancy management.
Profiling stem cells in differentiation
Mesenchymal stem cells, or MSCs, can differentiate into various cell types and have strong therapeutic potential for autoimmune disease and other disorders, but their development from embryonic stem cells remains poorly understood. In a recent paper published in the journal Molecular & Cellular Proteomics, a global research team led by Anja Billing of Weill Cornell Medicine-Qatar writes that they improved a protocol for developing MSCs from ES cells and then used this system to profile changes in the transcriptome, proteome and phosphoproteome during differentiation. The work identifies numerous regulatory changes that include changing expression of HOX transcription factors, long noncoding RNAs and signaling proteins. The multiomics approach also allowed the authors to appreciate, for example, that the proteins most phosphorylated are also the most strongly upregulated.
How protein stability shapes disease
Mutations in SECISBP2 — a protein necessary for selenoprotein synthesis — are linked to human disease, but patients with these mutations exhibit a high degree of phenotypic heterogeneity. To gain insight into how different mutations influence SECISBP2 function and clinical phenotypes, Wenchao Zhao of the Institut für Biochemie und Molekularbiologie and colleagues established two mouse models, each with a specific mutation that disrupts one of SECISBP2’s two selenoprotein-producing functions. In the Journal of Biological Chemistry, they write that they found that while one mutation nullified SECISBP2 function, the other mutation had tissue-dependent effects, suggesting that differing clinical phenotypes may be caused by varying SECISBP2 stability in different tissues.
Functions of a kidney regulator enzyme
Cytochromes CYP27B1 and CYP24A1 are known regulators of vitamin D in the kidney. However, the regulatory mechanism of the latter enzyme, which carries out separate functions in renal and nonrenal tissues, has not been clear. Mark B. Meyer, Seong Min Lee and colleagues in Wisconsin and Canada used ChIP-Seq and CRISPR/Cas9 genome editing to probe the genomic basis of CYP24A1 regulation in kidney and nonrenal target cells, or NRTCs. They uncovered regulatory regions downstream of the Cyp24a1 gene that, when specifically deleted, revealed a chromatin-based mechanism responsible for the CYP24A1’s differing functions in the kidney and NRTCs. Their study was published in the Journal of Biological Chemistry.
Lipid effects on endoplasmic reticulum structure
The endoplasmic reticulum, or ER, is a key organelle in all eukaryotic cells and plays essential roles in cell signaling and protein synthesis. The structure of the ER is integral to its function, and a number of human diseases including Alzheimer’s, Type 2 diabetes and certain cancers are associated with changes in that structure. Previous research primarily has focused on the role of proteins in maintaining ER structure, while the role of lipids (the main structural element of cell membranes) is less well studied.
A new paper by Gabriela Ulloa and an international team, published in the Journal of Lipid Research, highlights the important role that lipids play in mediating ER structure.
The researchers changed ER membrane lipid compositions in relatively inert sea urchin oocytes. By fluorescently labeling the ER membrane and using advanced microscopy techniques, they were able to follow changes in the proportion of two main types of ER structure: flat sheets and curved rods. Through this microscopy-based approach, diacylglycerol, or DAG, was identified as a key lipid in mediating ER structure, as the researchers found decreases in DAG skewed the rod-sheet split toward sheets.
Further experiments confirmed that the reintroduction of DAG recovered the normal balance, as did the addition of other lipids that energetically favor highly curved rodlike membranes, such as phosphatidylethanolamine. These ER structural modifications were independent of any changes in the number or type of proteins in the ER membrane, showing that the physical properties of the lipids themselves were driving the structural reformatting of the ER directly.
Beyond contributing to a deeper understanding of ER structural modifiers, this research could have applications in medicine. While most drugs target specific proteins, little drug development has targeted lipids directly to exert therapeutic benefit. With further study and more precise techniques, lipid-based targets for future therapies could emerge as a promising avenue in the treatment of ER-structure–associated human pathologies.
— Kian Kamgar–Parsi
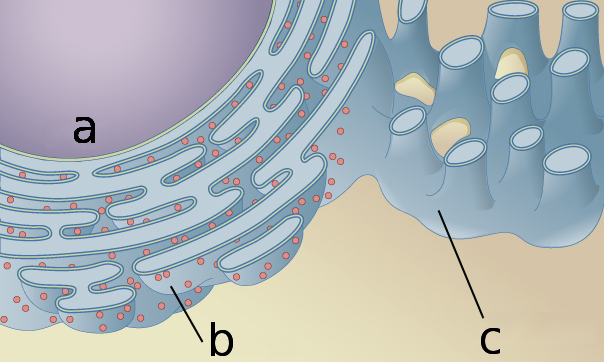 The endoplasmic reticulum surrounds the nucleus (a), and has a structure including both sheetlike (b) and rodlike (c) sections that contribute to its function.OpenStax/Wikimedia Commons
The endoplasmic reticulum surrounds the nucleus (a), and has a structure including both sheetlike (b) and rodlike (c) sections that contribute to its function.OpenStax/Wikimedia Commons
Sterol metabolism in the nematode
The metabolism of cholesterol is fundamental to eukaryotic life, as cholesterol contributes to hormone signaling, metabolism and maintaining cell structure. Given its importance in cellular function as well as its complexity, the cholesterol metabolism pathway is the subject of significant research to identify key divergences and elucidate major evolutionary points in eukaryotic life. Wenxu Zhou, Paxtyn Fisher and a team in W. David Nes’ lab at Texas Tech have studied the cholesterol metabolism pathways of nematode worms (a common, well-characterized test organism) to elucidate the specific processes therein.
In research published in the Journal of Lipid Research, the researchers used a variety of biochemical and spectroscopic tools to follow the evolution of cholesterol and sterol intermediates in the worms. They already knew that sterol metabolism occurs along two evolutionarily divergent branches in the worms, leading to either C4-methyl stanol or C4-methyl stenol products. Experiments in this study showed that the enzyme sterol C4alpha-methyltransferse was responsible for regulating sterol flux through these divergent pathways via a novel mechanism. This additional knowledge of the cholesterol pathway could provide insights into the evolutionary development of eukaryotic sterol metabolism.
Does this receptor really bind sugars?
Recent work has shown that a domain of the immunoglobulin E-binding receptor CD23 resembles a mammalian sugar-binding domain, but there have been conflicting reports regarding the receptor’s sugar-binding activity. Sabine A. F. Jégouzo, Hadar Feinberg and colleagues in the U.S. and U.K. used solid-phase binding competition assays, glycoprotein blotting experiments and glycan array analysis to investigate the binding activity of various mammalian CD23s, they write in the Journal of Biological Chemistry. While no sugar binding was detected for human CD23, a range of sugars were found to bind to cow and mouse CD23. The findings suggest that CD23 may play various roles across different species.
Glycan markers assigned with beam search
Many antibodies used to recognize stem cells recognize glycan epitopes on surface proteins or lipids. To understand these reagents better, an international team led by Nian Wu of Imperial College London used glycans synthesized in the lab to determine what these antibodies bind to with greatest efficacy. Using a computing approach called a beam search, which enables rapid searching through large data sets, in this case a glycan microarray, the authors resolve uncertainty about exactly what these antibodies bind to. The results, which were published in the journal Molecular & Cellular Proteomics, also shed light on stem cell glycobiology.
Breaking down bacterial biofilm adhesion
Cholera is caused by consuming water or food contaminated with the bacterium Vibrio cholerae.
Like nearly all other bacteria, Vibrio cholerae forms a biofilm — a bacterial community stitched together with a glue of polysaccharides, proteins and nucleic acids. It produces these biofilms in water while it awaits its thirsty host and then, once ingested, to protect against the acidic environment of the stomach, allowing it to survive and enter the small intestine, where it produces the toxin that causes severe diarrhea, dehydration and even death.
Becoming a biofilm has its benefits. Bunches of bacteria are less vulnerable than singletons to predation by protozoa and bacteriophages. They’re also more potent: Biofilms pack higher doses of bacteria and hyperinfective cells.
Scientists at Wesleyan University have been studying the bits and pieces of Vibrio cholerae biofilms found in water and the gut, and they recently reported a new finding about two components of the biofilm matrix in the Journal of Biological Chemistry.
Previous studies have suggested that the matrix proteins RbmC and Bap1 have redundant functions. When either is deleted, the biofilm grows as usual; when both are deleted, though, the cell clusters cannot attach to surfaces.
The study by Katherine Kaus and colleagues in Rich Olson’s lab at Wesleyan, however, suggests there may be nuanced, though important, differences in RbmC and Bap1.
The team found that Bap1 has a different binding activity than RbmC; Bap1 prefers anionic polysaccharides over complex N-glycans. These results indicate that Bap1 and RbmC may play critical but different roles in biofilm surface attachment.
“These studies in conjunction with structures of Bap1 complexed with carbohydrate ligands will provide a framework for understanding the network of complex molecular interactions that underlie biofilm assembly and adhesion in V. cholerae,” the authors wrote in JBC.
Kaus, now a postdoctoral researcher at the University of Texas Medical Branch in Galveston, said, “Our study is an exciting step forward in understanding the specific roles played by Bap1 and RbmC within the biofilm and how the slight differences in the two proteins might lead to important functional differences within the various environments encountered throughout the Vibrio cholerae lifecycle.”
— Angela Hopp
How strep prepares to infect
Streptococcus pyogenes — the bacterium responsible for strep throat and a diverse array of other diseases — relies on a surface polysaccharide called group A carbohydrate, or GAC, to infect human hosts. In a paper published in the Journal of Biological Chemistry, Azul Zorzoli and Benjamin H. Meyer of the University of Dundee and colleagues in Scotland and Russia write that they used molecular and synthetic biology approaches, biochemistry, radiolabeling and other techniques to show that the enzyme GacB performs a critical step in GAC biosynthesis. The results also indicated that the enzyme is conserved across the Streptococcus genus, including those with other types of surface carbohydrates, and may provide an opportunity to target early steps of GAC biosynthesis in S. pyogenes infections.
The upshot of blocking ubiquitination
Proteins modified by ubiquitin and small ubiquitinlike modifier, or SUMO, proteins are pivotal in myriad essential cellular processes and are elevated under proteotoxic conditions such as heat shock, but the reason why has not been clear. To understand better the basis for this effect, Zhe Sha and a team from Harvard Medical School and the Leiden University Medical Center blocked ubiquitination in human cell lines with a selective inhibitor of ubiquitin-activating enzyme. The treatment led to a large accumulation of SUMOylated proteins located in special nuclear bodies. These unexpected results, which were published in the Journal of Biological Chemistry, offer insight into the interplay between ubiquitination and SUMOylation under stress.
Elucidating a cervical cancer trigger
The degradation of tumor suppressor p53 by human papillomavirus, or HPV, oncoprotein E6 is a key step in the development of HPV-positive cervical cancer. E6 triggers the ubiquitination of p53 by associating with the ubiquitin ligase E6AP, but the function of the ligase is not well understood. By analyzing the p53 polyubiquitinated by E6AP in vitro, Yuji Masuda of Nagoya University and a team in Japan were able to elucidate details of the process. They found that p53 is multipolyubiquitinated with short chains and hypothesize that this is done in a stepwise manner. Together, the findings could inform the development of treatments for HPV-positive cervical cancer. The research was published in the Journal of Biological Chemistry.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.
The molecular orchestra of memory
Calcium, calmodulin and calcium/calmodulin-dependent kinase II form a molecular axis that turns fleeting neural activity into lasting memories. New research shows how memories are stabilized, and possibly even protected or repaired.

Differences in pili structure modulate bacterial behavior
Researchers demonstrate how small changes in the structure of hair-like protein appendages can affect the behavior of Acinetobacter bacteria.

Cholesterol regulatory genes predict liver transplant outcomes
Researchers identify a link between cholesterol-regulating genes and liver transplant success, which could improve donor screening and patient outcomes.





