Plasma membrane lipids mediate key interactions in assembly of filoviruses
Filoviruses, which include Ebola and Marburg, are lipid-enveloped viruses containing a negative-sense RNA genome encoding seven genes. While no Food and Drug Administration-approved drugs and vaccines exist for filoviruses, a recombinant Ebola vaccine, effective in animal models, appeared to help contain and limit the spread of a recent Ebola outbreak in the Democratic Republic of Congo.
One of the seven genes in Ebola and Marburg encodes the matrix protein known as VP40. VP40 is a peripheral protein that associates with the inner leaflet of the plasma membrane to regulate assembly of new virus particles. Strikingly, VP40 is able to form viruslike particles, or VLPs, from mammalian cells in the absence of other viral proteins. These particles are nearly indistinguishable from authentic virus and have provided researchers working outside of the highest biosafety-level labs with a system to investigate viral assembly and budding as well as the role of host lipids in these processes.
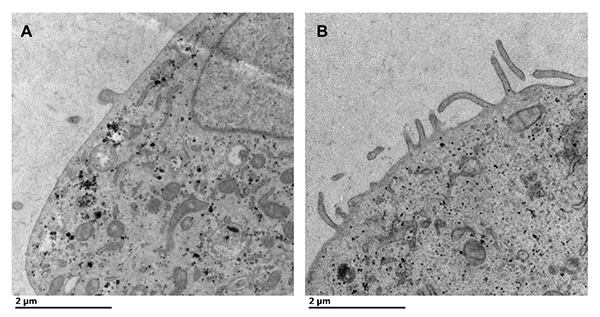 Transmission electron microscopy of cells expressing the Marburg matrix protein demonstrates viruslike particle formation from the cell plasma membrane. A: Mock transfected cells display few protrusions from the plasma membrane. B: Cells transfected with a Marburg matrix protein expression plasmid display remarkable viruslike particle formation from the plasma membrane of the cell after 24 hours.Courtesy of Monica Husby In research published in 2013, the X-ray structure of Ebola VP40, or eVP40, revealed a dimer mediated by a N-terminal domain alpha-helical interface (1). Mutation of the eVP40 dimer interface abrogated the ability of eVP40 to form VLPs and localize to the plasma membrane inner leaflet. The researchers discovered that eVP40 membrane binding regulated eVP40 oligomerization, as incubation of eVP40 with a membrane mimetic (dextran sulfate) led to formation of an eVP40 hexamer. Mutations in eVP40 that reduced hexamer formation abolished VLP formation.
Transmission electron microscopy of cells expressing the Marburg matrix protein demonstrates viruslike particle formation from the cell plasma membrane. A: Mock transfected cells display few protrusions from the plasma membrane. B: Cells transfected with a Marburg matrix protein expression plasmid display remarkable viruslike particle formation from the plasma membrane of the cell after 24 hours.Courtesy of Monica Husby In research published in 2013, the X-ray structure of Ebola VP40, or eVP40, revealed a dimer mediated by a N-terminal domain alpha-helical interface (1). Mutation of the eVP40 dimer interface abrogated the ability of eVP40 to form VLPs and localize to the plasma membrane inner leaflet. The researchers discovered that eVP40 membrane binding regulated eVP40 oligomerization, as incubation of eVP40 with a membrane mimetic (dextran sulfate) led to formation of an eVP40 hexamer. Mutations in eVP40 that reduced hexamer formation abolished VLP formation.
The same study found that eVP40 is a transformer protein with the ability to form an octameric ring structure that can bind RNA. The essential role of the eVP40 octamer in Ebola replication has not been resolved fully, but the octamer has not yet been observed at the plasma membrane inner leaflet or in VLPs or virions. Instead, recent research shows that the octamer plays a role in regulating viral transcription and is enriched in the host cell nucleus when compared to the eVP40 dimer (2). And while the role and significance of nuclear localization of eVP40 is still unknown, the eVP40 octamer binds plasma membrane lipids an order of magnitude less than the eVP40 dimer, which partially may explain its different cellular localization and role in viral replication.
We now know the role of plasma membrane lipids in Ebola virus assembly. Nearly two decades ago, experiments demonstrated that eVP40 bound vesicles containing phosphatidylserine, or PS; however, little information was available on lipid binding selectivity and the role of lipid binding in VLP formation. Now we know that eVP40 binds PS using two cationic loop regions in its C-terminal domain (2). PS binding by eVP40 is selective when compared to phosphatidylinositol, phosphatidylglycerol and phosphatidic acid. In support of the PS binding selectivity, several amino acids in the C-terminal domain contribute to PS recognition, including lysine, asparagine and serine residues. Swapping of native lysine for arginine (i.e., positive charge for positive charge) was not sufficient to restore PS selectivity.
Research published in 2015 showed that PS binding by eVP40 also was critical to eVP40 localization at the plasma membrane inner leaflet and VLP formation (3). In addition, eVP40 requires a significant pool of PS at the plasma membrane inner leaflet for efficient assembly and budding. A mutant cell line partially deficient in PS synthesis was critical in proposing this hypothesis, as VP40 assembly and budding was reduced significantly in mutant cells (with about a 30 percent reduction in PS levels) but restored upon adding PS back to these cells. PS binding also triggered conformational changes in eVP40 structure critical to appearance of VP40 hexamers and larger oligomers at the plasma membrane inner leaflet.
PS induces partial insertion of the eVP40 C-terminal domain into the hydrocarbon region of the plasma membrane, according to a 2013 research paper, which may aid in stabilizing the eVP40 oligomers that form and altering membrane structure important for viral budding.
For lipid researchers
Stay up to date on lipid research news and connect with other lipid scientists by joining the society’s lipid division. Learn more.
PS is not the only important plasma membrane lipid in EBOV assembly. A 2016 study found that phosphatidylinositol 4,5-bisphosphate, or PI(4,5)P2, modulates interactions of eVP40 oligomers with the membrane interface in vitro and in cell culture (4). In contrast to PS depletion, which significantly reduced eVP40 hexamer formation from dimers, PI(4,5)P2 depletion greatly reduced large eVP40 assemblies (those larger than hexamers and dodecamers) but didn’t significantly reduce eVP40 hexamers. Thus, PS and PI(4,5)P2 play contrasting roles in eVP40 assembly, with PS mediating the initial plasma membrane association and conformational change to form hexamers. Subsequently, PI(4,5)P2 stabilizes membrane interactions required to sustain large eVP40 oligomers to maintain viral budding.
Marburg VP40 has significantly different lipid binding properties from eVP40. A 2015 study showed that the mVP40 X-ray structure is conserved in overall structure with that of eVP40, harboring an N-terminal alpha-helical dimer interface and overall structural similarity between both the N- and C-terminal domains. However, the sequence similarity of the C-terminal domain in mVP40 was only 15 percent when compared to the eVP40 C-terminal domain. This difference in sequence is observed as amino acid changes in regions where VP40 associates with membranes. Marburg VP40 has a higher cationic charge density on its surface and does not have many of the key residues found interacting with PS in eVP40. Another study found that mVP40 behaves as an anionic charge sensor binding anionic lipids with promiscuity based upon the anionic charge density of the membrane interface. Neutralization of the plasma membrane interface in cell culture led to the rapid displacement of mVP40 from the plasma membrane, further supporting the anionic charge sensor hypothesis.
Host cell plasma membrane lipids are significant for more than filovirus assembly and budding. Research published in 2011 showed that budding. Ebola virus uses host PS to attach and enter cells through a process known as apoptotic mimicry. However, how PS levels on the outer membrane of the viral lipid envelope increased remains unclear. The plasma membrane of human cells is normally asymmetric, with PS concentrated in the inner leaflet. A clear connection to the role of PS in viral entry was made from the inner to outer plasma membrane leaflet when eVP40 is sufficient to induce PS exposure on the outer leaflet of the plasma membrane at sites of viral budding. A recent study found that a host scramblase, Xkr8, also contributes significantly to this process by helping the viral particles achieve a loss in plasma membrane PS asymmetry so that PS is available to interact with host cell receptors to mediate attachment and facilitate entry.
A challenge of studying lipids in filovirus replication has been the available viral systems to work with. Working with live virus requires the highest biosafety-level facilities, and the process of virus neutralization affects lipid measurements, limiting the studies performed on virus-induced lipid metabolic changes. However, lower biosafety-level systems are available that allow for a more complete representation of the viral life cycle. For instance, a 2014 study found that a tetracistronic minigenome system allows for all seven Ebola virus proteins to be expressed in human cells, allowing budding and viral entry to be studied in more complete fashion. Implementation of such assay systems or generation of new assay systems should allow for mechanistic investigation of lipid metabolic changes in cells infected by filoviruses, perhaps unveiling new drug targets or biomarkers of disease in the viral life cycle.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

