A simple method to determine phase preference of proteins on live cell membranes
Scientists at National University of Singapore have demonstrated a simple and fast method to determine if a biomolecule partitions into lipid domains on live cell membranes. Their work was published in the Journal of Lipid Research.
Cellular membranes are dynamic assemblies of lipids and proteins with some components organized as domains. Proper cell function requires the partitioning of lipids and proteins into these domains, which are often rich in cholesterol and sphingolipids. However, they are too small (10-200 nm in size) and dynamic (possibly only tens of milliseconds in lifetime) to be observed even with modern super-resolution microscopy techniques.
Traditional methods to determine domain localization involve biochemical assays that require many cells, are prone to artifacts as they are conducted in vitro, and are slow. Although fluorescence-based techniques can probe these domains in live cells, they require specialized instrumentation and are often difficult to interpret.
The research team at NUS developed a simple fluidizer-based method to determine if a molecule prefers to partition into lipid domains on cell membranes.
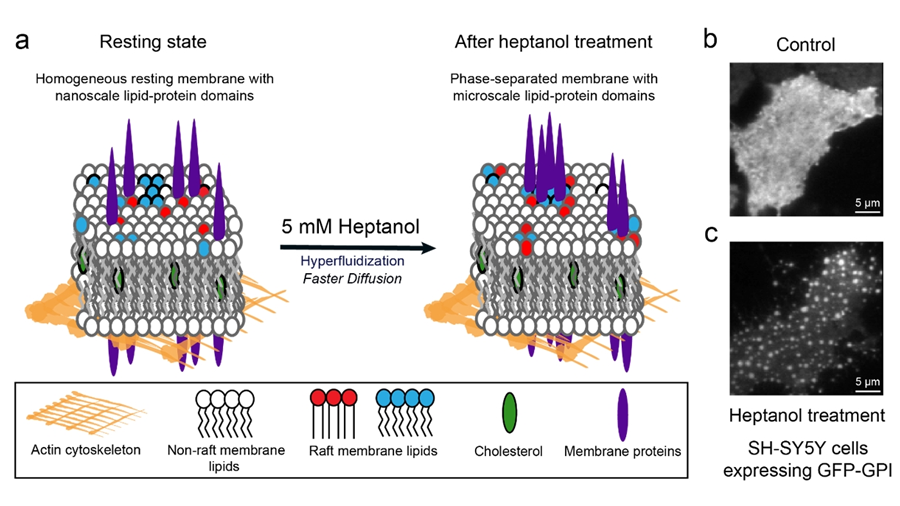
The team added heptanol to live cells and showed that within 15 minutes it induces clustering of the nanometer-size lipid domains into larger micrometer-size domains that are easily detectable by standard fluorescence microscopes. The method works with both molecules that are genetically labelled with fluorescent proteins and those labelled using extrinsic labels, for example, antibodies.
The work was conducted in the lab of Thorsten Wohland and led by first author Anjali Gupta, who is now a research fellow at Harvard Medical School and Boston Children's Hospital.
Gupta explained the significance of studying phase preference of molecules in membranes.
"Phase preference of molecules in membranes is fundamentally crucial for the essential biological processes originating at membranes, such as T-cell activation, a critical step during an immune response," she said. "Knowledge of the phase preference of molecules will support therapeutic development based on the modulation of lipid domains."
Wohland said: “The phase preference of molecules used to be difficult and time-consuming to establish. This new method, detected by chance, provides results in at most 15 minutes on live cells and can essentially be seen by eye in a simple microscope.”
The team hopes that this technique will enable a quick and facile identification of domain localization and will aid the wider research community.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

