Researchers target cell membrane for cancer research
Robert Chapkin has spent decades studying the molecular roles components of nutrition play in protein signaling and the prevention of diseases. His lab's recent discoveries about lipids and the cell membrane could revolutionize translational cancer research.
"Cell membranes are the lipid environment in which many proteins function," said Chapkin, a professor of nutrition at Texas A&M University. "It is now appreciated that protein and lipids assemble to form distinct micro- or nanodomains (clusters) that facilitate key signaling events."
Cell membrane composition is altered in diseases such as cancer and obesity. Chapkin believes that membrane therapy — the modulation of cellular membrane lipid composition and organization — might be an effective therapeutic strategy.
"The central idea is that if you alter the composition of the cell membrane, you can potentially alter the functionality of the proteins within the membrane and thus the disease overall," he said.
More than a decade ago, Chapkin's lab discovered that docosahexaenoic acid, or DHA, a well-known dietary omega-3 fatty acid and chemoprotectant, suppresses the functionality of epidermal growth factor receptor, or EGFR, a protein in the cell membrane that drives the formation of many types of cancer, including colon cancer.
But how does DHA suppress the function of the EGFR protein? Natividad "Robert" Fuentes, a former graduate student in the Chapkin lab and first author on the lab's recent paper in the Journal of Lipid Research, uncovered some groundbreaking molecular insights into this mechanism. Using cell models and animal models and a cutting-edge technique called super-resolution microscopy, he studied the changes to the lipid membrane and EGFR after DHA incorporation.
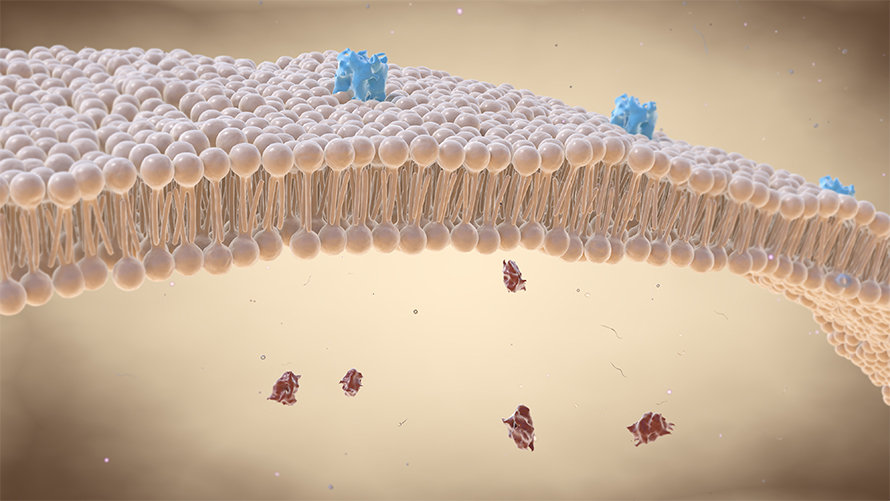
"We found that when DHA is incorporated, it alters the localization of the lipid bilayer with EGFR," Fuentes said. "It alters the spatial orientation of the protein in the lipid bilayer."
Why does this matter? It turns out that the architecture of the protein within the lipid bilayer of the cell membrane is one of the factors that drives its function. This might explain why DHA incorporation suppresses EGFR signaling.
Fuentes said he believes such membrane therapy could synergize with other cancer treatments. "The fatty acids that modulate the lipid bilayer are completely innocuous to humans and could potentially be used as adjuvants to suppress the functionality of proteins that drive cancer."
As a postdoc at the University of Texas MD Anderson Cancer Center, Fuentes now uses membrane therapy in translational pancreatic cancer research.
"Pancreatic cancer is resistant to many therapies," he said. "Part of my work is to study how disrupting the pancreatic cell membrane might improve the efficacy of cancer therapeutics."
With membrane therapy still in its infancy, Fuentes believes it will be applicable in other research areas. "Membrane therapy holds promise for any disease states where receptor clustering within the cell membrane is affected," he said. "For instance, it could be used in diabetes research to target the insulin receptor and insulin signaling."
Chapkin is eager to explore the more mechanistic nuances and specificity of membrane therapy and study other potential players.
"We will be researching other preventative components of nutrition and target proteins," he said. "There is so much exciting work to be done in this field."
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

