Innovative platform empowers scientists to transform venoms into therapeutics
Some of the most successful and effective drugs, like Ozempic, come from animal venoms. However, scientists usually only discover the therapeutic potential of venoms by chance. Recently, an international group of researchers, led by Meng-Hsuan Hsiao of Ben Larman ’s laboratory at Johns Hopkins University, set out to change that by developing a workflow to accelerate drug discovery from peptides, or short sequences of amino acids, with sequences and structures that resemble animal venoms. They published their study in Molecular & Cellular Proteomics.
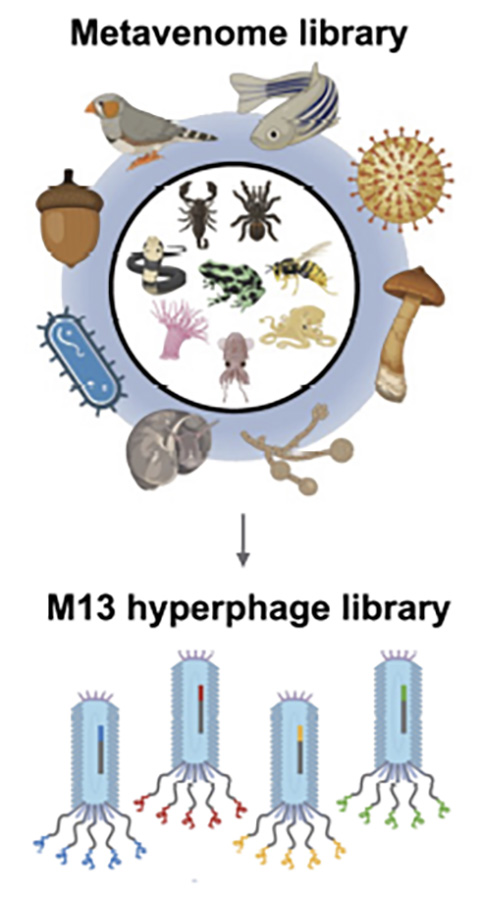
Hsiao and colleagues demonstrated that their platform could evaluate over 10,000 venom-like peptides’ ability to bind specific receptors on human cells. Larman’s group teamed up with Martin Steinegger, an assistant professor at Seoul National University, who built a “metavenome library” from known animal venom sequences and additional peptides based on sequence homology.
“A library allows us to explore a much larger space of possible lead compounds for drugs,” Larman said. “You don’t even need to have a starting hypothesis.”
The researchers expressed peptides from this library on the surface of bacteriophages, or bacteria-infecting viruses, a technique also known as phage display. For their platform, Larman’s team used what he called a specific “flavor” of phage: the M13 hyperphage.
Hyperphages carry genetic mutations that make them display five copies of venom-like peptides on their surface. Therefore, the displayed peptides could interact with multiple receptors, amplifying the signal and allowing the researchers to detect weaker interactions.
“The way we used hyperphages to display venoms has never been done before,” said Larman.
Unlike other phages, M13 phages are secreted through the bacterial periplasm, the space between the bacteria’s inner and outer cell membranes. The periplasm has an oxidizing chemical environment, Larman explained, and this helps preserve the venom peptides’ disulfide bonds, which are critical for their 3D structures. These disulfide bonds make venoms attractive therapeutics because they are highly compact and resistant to degradation.
Using their platform, Larman’s team identified six proteins in the metavenome library that bind and activate the human itch receptor MAS-related G protein-coupled receptor X4, or MRGPRX4. They found that these proteins share a unique folding pattern called the Kunitz domain, which is commonly known to inhibit serine protease activity. In the future, Larman’s team plans to expand their metavenome library to include even more peptides in search of a potent MRGPRX4 antagonist that can be developed into an anti-itch drug.
“I think that if you can find an agonist, you can also find an antagonist,” Larman said, “And we can likely achieve this by expanding our library.”
Larman said he hopes to integrate generative artificial intelligence into their system to broaden the library to include novel venom-like peptides.
“Once set up and integrated,” he said. “I can see this cycle of computational prediction and experimental validation being really powerful in drug discovery.”
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.
The molecular orchestra of memory
Calcium, calmodulin and calcium/calmodulin-dependent kinase II form a molecular axis that turns fleeting neural activity into lasting memories. New research shows how memories are stabilized, and possibly even protected or repaired.

