From the journals: MCP
A new way to label proteins in the brain. Novel targets for a ubiquitination enzyme. Ultrasensitive tools detect extracellular vesicles. Read about papers on these topics recently published in Molecular & Cellular Proteomics.
A new way to label proteins in the brain
Several methods exist to isolate and study proteins and their interactions; however, studying changes in proteins across different cell types in the brain can be a challenge because most brain cells do not survive the harsh isolation process.
TurboID is a new and efficient biotin ligase that can label proteins in cells within a radius of 10 nanometers. It is nontoxic and fast, generating labeling within 10 minutes instead of the 18 hours required by the biotin ligase BioID. With this powerful tool, researchers can analyze the global proteome within specific cell types in multicellular models, in both homeostatic and diseased states.
TurboID has been applied in proteomic studies of different cell types in animal models, although it is important to determine whether the protein profile captured by the TurboID method is indeed representative of the whole cell in both resting and perturbed states, can differentiate between cell types, and whether TurboID overexpression and biotin labeling of proteins affects cellular processes.
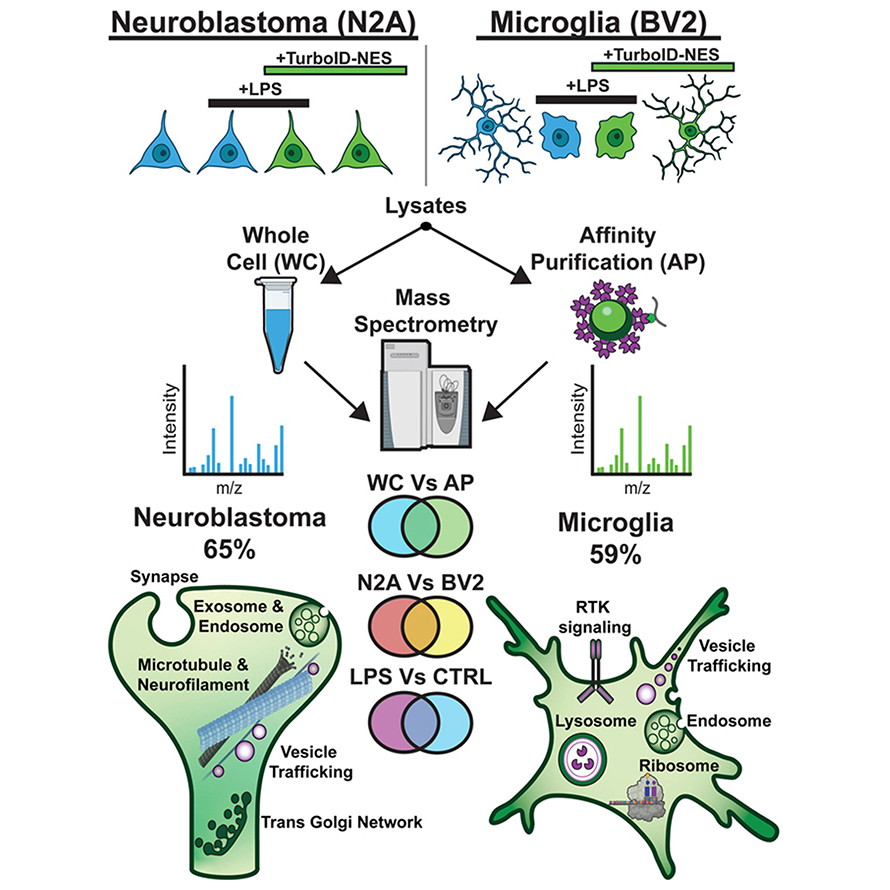
Sydney Sunna and colleagues at Emory University directed TurboID to label proteins in the cytosol of microglial and neuronal cells, using a nuclear export sequence. They hypothesized that TurboID can label enough proteins to distinguish between the cell types and that the labeled proteins include proteins known to respond to an inflammatory stimulus without toxicity.
In the journal Molecular & Cellular Proteomics, they report that TurboID can label more than 50% of the whole proteome in each cell type and identify several neuron-enriched and microglial-enriched proteins; thus, distinguishing the two cell types. Overexpression of TurboID did not impair any metabolic function. This method can also capture changes in microglia in response to inflammatory stimulation.
Many proteins identified by this method are implicated in neurodegenerative diseases such as Parkinson’s, Alzheimer’s and amyotrophic lateral sclerosis. Moreover, TurboID may be able to investigate the mislocalization of proteins from the nucleus to the cytosol, which is known to occur in these diseases. These in vitro studies lay the groundwork for ongoing and future applications of TurboID to explore responses of different brain cell types in mouse models of diseases involving neurodegeneration and inflammation.
Novel targets for a ubiquitination enzyme
Ubiquitination regulates various processes in cells, such as protein homeostasis and the removal of unwanted proteins. A trio of enzymes carries out this posttranslational modification: ubiquitin activating enzymes, or E1; ubiquitin-conjugating enzymes, or E2; and ubiquitin ligases, or E3. These enzymes attach a protein called ubiquitin to a target protein, labeling it for consequences such as protein degradation, change in protein activity or cellular localization. Dysregulation of ubiquitin–enzyme machinery is seen in many diseases including cancer.
Of the trio, the target proteins for E2 have been the most elusive; these enzymes have multiple targets and the enzyme–substrate interactions are transient. Zeliha Yalcin and colleagues at the Netherlands Cancer Institute discovered the protein targets for one particular E2 enzyme, UBE2D3, which has shown promiscuous activity in in vitroexperiments in test tubes, but researchers know less about what it does in organisms.
In a recent article in Molecular & Cellular Proteomics, the team reports they identified UBE2D3 substrates using a combination of proteomics techniques. They studied the effect of depleting UBE2D3 on the abundance and ubiquitination of proteins in the cell and found that several proteins involved in retinol metabolism or cell adhesion had higher or lower abundance, specifically CRABP1 and TSPAN8. Using a different technique, they also confirmed two protein substrates of UBE2D3 that are involved in regulating ribosome-associated protein quality control.
These results will inform future research on the cellular mechanisms of ubiquitination and cancer.
Ultrasensitive tools detect extracellular vesicles
Extracellular vesicles, or EVs, are heterogeneous, phospholipid-rich particles secreted by cells that carry biomolecules such as proteins, nucleic acids and metabolites. The discovery that EVs facilitate long-distance communication between cells has piqued researchers’ interest. Both healthy and diseased cells secrete EVs, and their internal contents change depending on the disease state. Thus, clinicians can use them to diagnose and monitor diseases including neurodegenerative conditions such as Parkinson’s.
However, researchers have difficulty finding appropriate protein biomarkers to selectively isolate cell-type–specific EVs due to the variety of internal and external proteins and differences in sizes as well as a low abundance of EVs in biofluids such as blood and cerebrospinal fluid.
In a recent article in Molecular & Cellular Proteomics, Adnan Shami-Shah and colleagues at Harvard Medical School report on ultrasensitive protein detection technologies that could help detect rare EVs. These include technologies with high sensitivity such as Luminex, best known for multiplexing 500 analytes that require a sub-microliter volume capable of detecting multiple surface proteins on EVs, and SOMAscan, which can quantitatively analyze 7,000 proteins concurrently with a high femtomolar sensitivity and a wide dynamic range.
Among other existing ultrasensitive technologies, Meso-Scale Discovery has a robust biofluids matrix tolerance with high detection sensitivity and specificity through its capacity to minimize autofluorescence background signal. Proximity extension assay can process 3,072 analytes and has high specificity due to a unique oligonucleotide hybridization step. Droplet-based EV analysis has a high sampling efficiency (about 20 million droplets per minute) and a limit of detection of 9 EVs/µL. Single-molecule array (Simoa for short) produces a localized signal to detect rare EV proteins and can switch from digital to analog reading within the same assay to detect varying abundance of EV subpopulations. Last but not least, molecular on-bead signal amplification for individual counting (MOSAIC for short) can detect rare EV proteins through sub-attomolar sensitivities and an enhanced noise-to-signal ratio.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

