Variety is not a spice for some lipids!
We all learn that phospholipids are composed of glycerol, to which fatty acids and a defining head group are attached. We also know that many phospholipids have a characteristic set of attached fatty acids. Why such specificity exists and which physiological roles they play has eluded us. In this month’s Lipid News article, Richard Epand of McMaster University describes his recent work that uncovered how this enrichment is achieved for the important signaling lipids called phosphoinositides.
Fatty-acid composition in phospholipids sometimes happens after their synthesis, at which point the attached fatty acids are exchanged for other fatty acids in a process known as remodeling. For the phosphoinositides, the selectivity occurs not only via remodeling after the synthesis of the phospholipid but also during their synthesis. These phospholipids are enriched heavily with a select set of fatty acids called stearic acid and arachidonic acid. Epand’s lab showed that two important enzymes involved in their synthesis, diacylglycerol kinase-epsilon and cytidine diphosphate synthease-2, prefer precursor substrates with the selected fatty acids. Epand’s article describes new findings regarding how fatty-acid selectivity is achieved for other lipids as well.
These studies offer us a glimpse of the mechanisms of fatty-acid specificity within lipid classes. Why is this important? What role do select fatty acids play in physiological processes? We now may be in a better position to start answering these questions. – Daniel Raben
Enrichment of acyl chains in the lipids of the phosphatidylinositol cycle
By Richard Epand
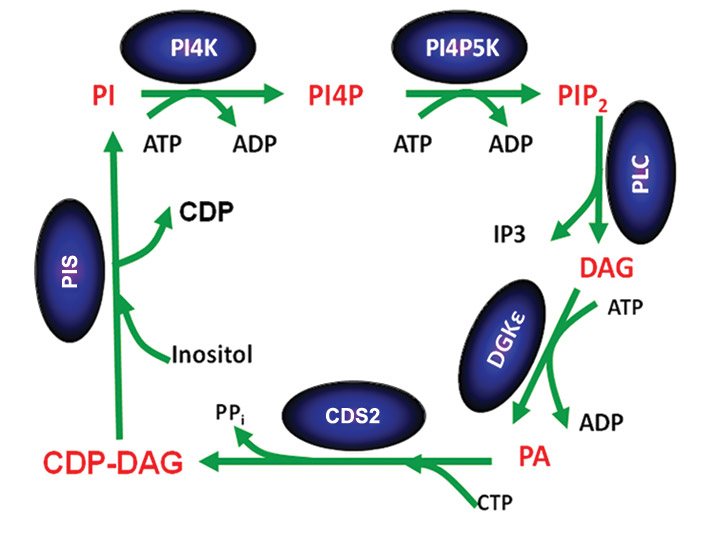
Phosphorylated forms of phosphatidylinositol play important roles in signaling and cell function. It has been known for many years that these lipids are synthesized in the phosphatidylinositol cycle and are highly enriched with 1-stearoyl-2-arachidonoyl species. Several mechanisms contribute to this enrichment including the remodeling of synthesized phospholipids through the action of acyl transferases (the Lands pathway) as well as through the specificity of the enzymes of the phosphatidylinositol cycle (see figure).
In the phosphatidylinositol cycle, a diacylglycerol kinase phosphorylates diacylglycerol. Although there are 10 different diacylglycerol kinase isoforms in mammals, in addition to gene-splice variants, one isoform called DGKε shows specificity for diacylglycerol species that reflect the acyl chains found in phosphatidylinositol. The phosphatidic acid product of DGKε catalysis is then converted to CDP-diacylglycerol (CDP-DAG), catalyzed by the enzyme CDP-DAG synthase, known as CDS.
CDS2, one of the two isoforms of mammalian CDS, also exhibits acyl-chain specificity for its substrate phosphatidic acid, favoring 1-stearoyl-2-arachidonoyl species, like DGKε
It is important that the phosphatidylinositol cycle describes a cyclical metabolic pathway so that the intermediates of the cycle are regenerated each time the cycle goes around. As a consequence, the preferential incorporation of particular acyl chains at one step in the cycle would be magnified because of progressive enrichment in each cycle. The lipid intermediates of the cycle have to be segregated from other forms in the cell, because most species of these intermediates would not be enriched with 1-stearoyl-2-arachidonoyl species. This segregation is difficult to achieve, because different steps of the cycle occur in two different membranes: the plasma membrane and the endoplasmic reticulum. This is a current area of research.
Once phosphatidylinositol is formed, the acyl-chain enrichment with 1-stearoyl-2-arachidonoyl species does not change much in forming PI4P and PIP2 (see figure). In addition, we find that conversion of CDP-DAG to phosphatidylinositol, catalyzed by phosphatidylinositol synthase, or PIS for short, does not result in acyl-chain enrichment. Therefore, only two enzymes – DGKε and CDS2 – are principally responsible for acyl-chain enrichment. We suggest that the extent of acyl-chain enrichment with 1-stearoyl-2-arachidonoyl species depends to a large extent on the level of expression of these isoforms in different organs.
Interestingly, the enzyme PIS, which does not exhibit acyl-chain preference, is expressed in only one form. In contrast, there are multiple isoforms for DGK and CDS, only one of which shows acyl-chain specificity. Hence, when the availability of arachidonic acid, an essential fatty acid, is low or the expression of DGKε or CDS2 is low, phosphatidylinositol synthesis still can continue with the substitution of other isoforms of DGK or CDS.
Other organisms have phosphatidylinositol species that are different from the 1-stearoyl-2-arachidonoyl forms found in mammals. Plants have mostly linoleic acid at the sn-2 position. The two major phosphatidylinositol species in yeast are 34:1 and 32:1, making up 56 percent of the phosphatidylinositol In the case of the Amoebozoa, Dictyostelium discoideum, the major phosphatidylinositol species, is an ether-linked lipid 1-hexadecyl-2-(11Z-octadecanoyl)-sn-glycero-3-phospho-(1’-myo-inositol). It appears that the phosphatidylinositol of Dictyostelium discoideum is not formed from diacylglycerol through the phosphatidylinositol cycle but rather from phosphatidic acid.
All eukaryotes have only one or a few species of phosphatidylinositol with particular hydrocarbon chains. This observation suggests that the hydrocarbon chains of phosphatidylinositol and its phosphorylated species are important to their functions. Specific acyl chains on other lipids also may be required for specific functions. We are just beginning to unravel the mechanisms by which hydrocarbon chain specificity determines physiological function.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.


