The wild frontier of model organism research
In 2006, Wojciech Pisula snuck behind a gas station in Warsaw, Poland to catch a rat. A psychologist at the Polish Academy of Sciences, Pisula had been sending lab rats through mazes for more than a decade and was convinced that these mild animals were no longer representative of their kind. Through hundreds of generations, he knew, the rats — most often Rattus norvegicus, the Norway rat — had, like other laboratory organisms, been domesticated and bred by scientists to select for traits that made them easier to study. To understand just how different his lab rats had become from their wild relatives, Pisula decided to bring some wild rats into captivity. He set traps by the dumpsters here and at other locations around the city and waited.
processes in many fields of research. But until recently, almost nothing
was known about the worm’s life in the wild.
Over the next 15 years, Pisula and a group of psychologists studied rats from the Warsaw wilds. With new captures introduced periodically, the scientists observed clear differences between the wild and the lab strains — especially the albino rats important to medical research. For instance, where lab rats relied almost entirely on their sense of smell and long whiskers to navigate the world, the wild rats relied on their eyes; they froze in alarm unless Pisula watched them in the dark. Some of these differences started to disappear after just three generations in the lab. “It is a change,” said Pisula.
Last year, Pisula stopped the experiment. The wild rats were too stressed living in the lab and maintaining them had become too expensive and difficult. But he had shown what he set out to: One could learn something new by studying the wild counterparts of one of biology’s classic model organisms — the select few animals, plants, and microbes that scientists rely on to conduct laboratory experiments investigating how life works.
From the fruit fly to brewer’s yeast to the proverbial lab rat, model organisms have been behind many of modern biology’s basic discoveries and continue to dominate research today. But as with the rat, research on these classic models has largely focused on domesticated strains studied in reductive laboratory settings. Despite the scrutiny in the lab, for a long time relatively little was known about their lives in the wild.
Scientists like Pisula have sought to change that, expanding the scope of research on the classic strains by incorporating more natural diversity in experiments and considering their ecology and evolutionary history. Researchers say this approach is not likely to lead to the kinds of fundamental discoveries made in the classic models. But wild counterparts have offered a fresh look at connections across all levels of biological complexity, from the ecosystem to the cell — and have even, in some cases, led to findings relevant to human health, from Alzheimer’s disease to cancer treatments. Ian Baldwin, a chemical ecologist at the Max Planck Institute for Chemical Ecology in Germany and editor of an ongoing series in the journal eLife that captures some of the results of this wide-ranging work, calls it the “wild frontier” of model organism research.
“We realized that the models are all really weird,” said Baldwin. “They’ve all got traits that made them very easy to be domesticated in the laboratory. And some of the nearest relatives to these models are completely different.”
Learning “a lot of new biology,” he added, was as simple as looking beyond the lab.
While anything could be a model organism in theory, most biological research and funding has focused on just a few species. This narrow approach has had stunning results, enabling scientists to learn about biological processes shared by all, or most, living things. In the 1850s, the microbiologist Louis Pasteur used the yeast Saccharomyces cerevisiae to outline how cells produce energy with and without oxygen. In the early 20th century, Thomas Hunt Morgan used the fruit fly Drosophila melanogaster to understand aspects of biological inheritance. In the early 1960s, Francis Crick and others used the bacterium Escherichia coli to crack the genetic code. Today, the house mouse, and to a lesser extent the Norway rat, are the workhorses of medical research, and the mustard plant Arabidopsis thaliana is at the root of plant genetics and associated advances in agriculture. These and a few others have been compared to the United Nations Security Council: the preeminent models among the general assembly of life.
But while much is known about the biology of the classic model organisms, their lives outside the lab — their natural histories — are less understood. And what research has been carried out on their ecology, evolution, and natural variation is rarely incorporated in laboratory experiments focused on basic processes at the molecular or cellular level. Many scientists who have studied these model organisms tended to treat them as a step in a chemical reaction, said Baldwin, and “hadn’t really thought much about the organism as an organism.”
Sarah Farris, a neurobiologist at West Virginia University who recently wrote about the need to support more research beyond the classic models, cited a colleague’s analogy of using a zoom lens on a camera. “These genetic model systems provide an extremely high-resolution picture, but a very narrow field of view,” she said. “Because you're looking at just one organism, you have really no idea of the context of that genetic pathway, or the function of that structure outside of this animal living in a lab.”
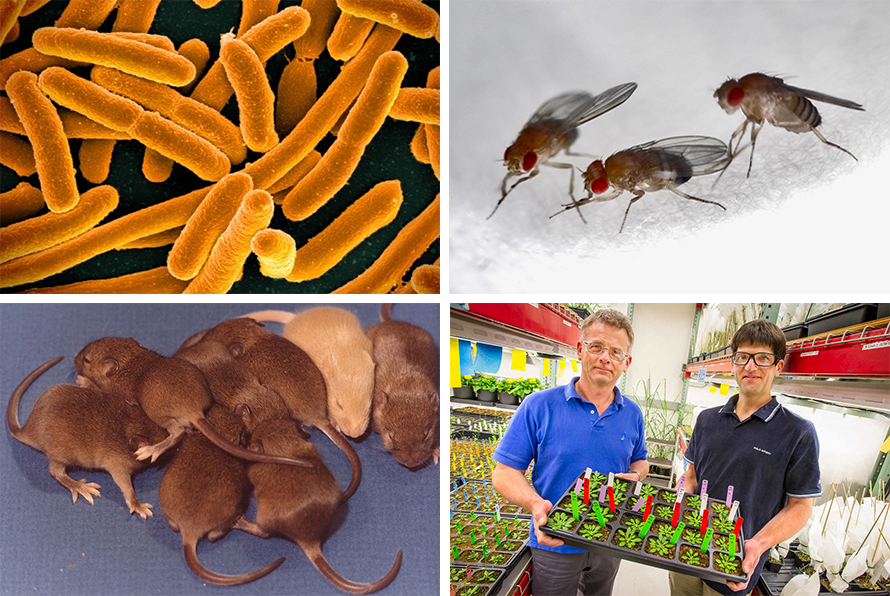

Through evolution, plants, animals, and microbes are shaped by their ability to live in particular environments, giving rise to the diversity of life across the planet. The classic model organisms, though they are as much a part of this diversity as wolves or octopuses, aren’t often put within this broader view. The problem with this approach, at least in the eyes of scientists like Baldwin and Farris, is that it overlooks the reasons why a creature is the way it is. It’s like knowing everything there is to know about a hammer, except what it’s used for.
That narrow view is expanding. Powerful genetic tools have become cheap and agile enough for biologists to apply them beyond the laboratory strains of the classic models. Just a few years ago, it was a luxury to have decoded any organism’s entire set of genetic material, its genome. Today, DNA sequencing is fast and cheap enough to allow whole-genome sequencing of entire populations, allowing an unprecedented view of genetic diversity and its consequences. Similarly, editing genes in order to study their function used to be possible only with complex processes developed for just a few organisms; now, gene-editing tools like CRISPR/Cas9 promise to enable scientists to alter the genes of pretty much any living thing, from honeybees to squid. “All of the interesting things that have evolved on this planet in the past 40 million years of biological evolution are now on the potential radar,” said Baldwin. The classic models, with decades of laboratory scrutiny to back them up, just have a head start.
Take the worm Caenorhabditis elegans, a tiny, transparent nematode used to model biological processes in many fields of research, including for human disease. In the past, almost all research on the worm was conducted on a single strain isolated from a compost pile in Bristol, England, known as the N2 lineage. Biologist Sydney Brenner and others launched C. elegans to model organism fame in the 1960s and 1970s, using it, among other things, to investigate how genes control how organisms develop. Accepting a Nobel Prize in 2002 for these efforts, Brenner said the worm should be receiving the prize along with him. But despite all the attention, for a century, almost nothing was known about C. elegans’ life in the wild that wasn’t inferred from the lab. “I mean, for so long, we didn’t know where C. elegans lived,” said Baldwin. Nor did scientists know what the worms ate. Most scientists fed the worms E. coli simply because there were lots of the bacteria lying around.
Over the past 20 years, however, biologists have filled in the picture, studying wild populations of C. elegans and other related species. In one paper in the eLife series, Lise Frézal and Marie-Anne Félix of the National Center for Scientific Research in Paris explain how C. elegans lead boom-and-bust lives, with their populations spiking each fall before crashing each winter (at least in Europe). The worms make their home in rotting plant matter, sometimes associated with human crops and compost heaps. And unlike in the lab, wild worms live together with a cosmopolitan set of microbes, arthropods, and other invertebrates.
Erik Andersen, a geneticist at Northwestern University, is among the researchers using this burgeoning natural history of the worm to study evolution and the consequences of natural variation for worms, and indirectly, for humans. His research focuses on a database of worm diversity he started building in 2014 after a study he co-authored on 200 wild strains of C. elegans provoked a flurry of interest from other researchers. While his team collected worms from Illinois to Hawaii, other colleagues and citizen scientists mailed in samples from all over the world.
In 2001, there were just 36 strains of C. elegans available for research. Today, frozen in Andersen’s repository there are more than 1,000 strains of C. elegans and more than 2,000 from two related species, all with ecological data and complete DNA sequences, and all available for free. Not only is it the most diverse collection of these creatures in the world, Andersen said — it is the most diverse collection within a genus of any living thing.
Andersen uses this natural variation to understand the genes involved in complex traits in the worm. By comparing wild strains, he can map differences in things like body size and behavior to genetic differences. These connections help to identify the function of particular genes, linking an organism’s genetic code — its genotype — with the observable expression of that code — its phenotype. Andersen also keeps track of where each worm was collected to understand how the variation among the worms may have helped them adapt to particular environments. These links across levels of biological complexity offer insight into deep processes of evolution, such as the origins of C. elegans’ distinctive sex lives — most worms are hermaphrodites which can either fertilize themselves or mate with the rare male that comes along — or how worms across the globe have adapted to human activity over the past two centuries.
“What makes organisms the way they are in a particular environment, wherever it might be?” asked Andersen. “That's the type of question we can now do.” No longer restricted to familiar worms in petri dishes and test tubes, “it can actually be going out there physically handling organisms,” Andersen continued, “and then going back to the lab, moving clear liquids from tube to tube and making connections that hadn't been possible previously.”
Though making direct connections between this work in worms and humans is difficult (and not the primary aim of this basic research), said Andersen, it is possible. In 2017, for instance, he and his colleagues looked at how worms responded to a toxin used in chemotherapy called etoposide. Had they used just laboratory strains, all worms would have seemed to respond the same way, but by comparing worms from different wild populations, the team showed how sensitivity to the toxin varied depending on the strain. They were then able to identify the genetic basis for those different sensitivities, narrowing it down to a single gene that encodes a protein involved in unwinding DNA. Because variation in residues from the corresponding proteins in humans determines sensitivity to etoposide, the finding in worms allowed Andersen’s group to explain why different parts of the human body are more or less sensitive to the toxin — why etoposide might harm a patient’s heart, but not their lungs. Without the wild worms, this insight would have been missed.
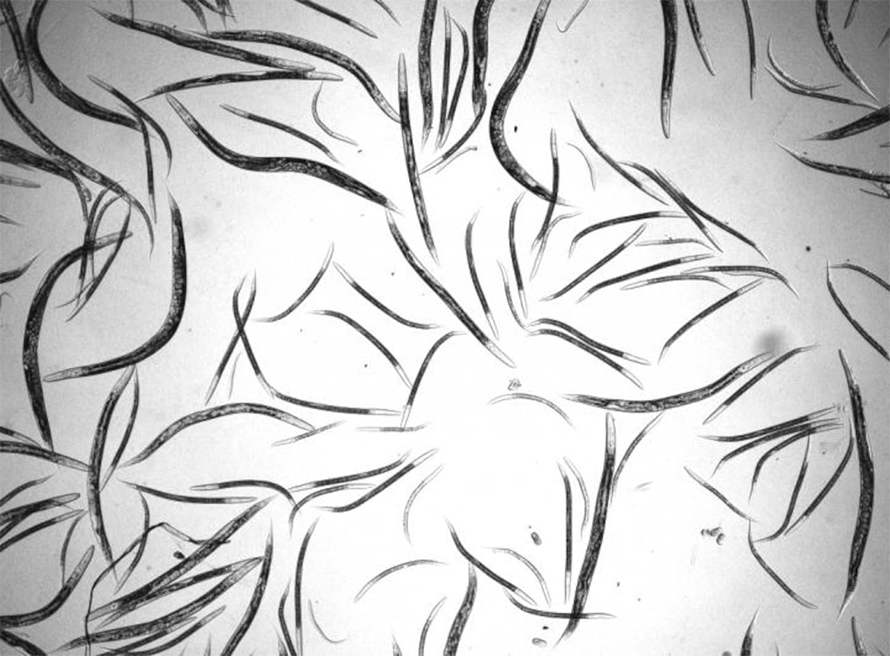

As with the worm, biologists have increasingly looked to natural history and diversity in other classic models. For instance, Megan Phifer-Rixey, a biologist at Monmouth University in New Jersey, uses wild house mice to study evolution. Though the inbred laboratory strains used in most research were “really powerful,” said Phifer-Rixey, “they didn't actually represent the full scope of variation that was present in the wild.” Collecting wild mice from houses and horse barns, she and her colleagues have amassed mice from the nearby Jersey Shore to Florida, and people have sent her animals from as far away as Iran. She and others have used the wild mice to discover more about the ubiquitous critters — for example, she and her colleagues found that the microbes in their gut are primarily transmitted from mother to offspring, rather than from living in close proximity with each other.
“People are excited and interested in that work and figuring out how it's relevant to their more clinical work,” said Phifer-Rixey. At the Jackson Laboratory in Maine, for instance, researchers have used wild-derived mice to study how genetic variation affects cells implicated in Alzheimer’s disease. Another group of researchers inserted embryos from laboratory mice into wild mice. Inoculated with the mothers’ wild microorganisms during birth, the offspring ended up with more diverse microorganisms in their gut. This could make the hybrid mice — “wildlings,” as the researchers called them — better models for testing drugs that impact the human immune system, which is influenced by the diversity of the gut microbiome.
“It's really a two-way street,” said Phifer-Rixey. “The wild mice have a lot to offer our studies of laboratory mice and medicine and genetics.” But laboratory work also informs the research on wild mice. “When I'm asking a question about what's happening in the wild,” she added, “it sure helps to have a genome sequenced and to know what the genes in the genome do.”
Research on the fruit fly Drosophila melanogaster has also taken a wild turn. For a century, laboratory populations of this single species and its laboratory mutants enabled key advances in genetics, including as one of the first multicellular organisms sequenced by scientists in 2000 — C. elegans was the very first — paving the way for the completion of the vastly larger human genome. As sequencing has become faster and cheaper, much research on the fruit fly has moved beyond the familiar laboratory strains. This year an international collaboration of researchers unveiled a new dataset containing the genomes of 271 strains of D. melanogaster collected from more than 20 countries on four continents. “That’s just really an amazing advance forward that we didn't have just a year ago,” said Patrick O’Grady, an evolutionary biologist at Cornell University and director of the National Drosophila Species Stock Center. “People look at stuff from all over the world now.”
And research on S. cerevisiae, perhaps the most well-characterized of all model organisms, has seen a similar expansion. Though early research on the fungi involved genetic variation — after all, artisans use it to make wine, beer, and sake, each of which is associated with different yeast breeds — most yeast biologists eventually narrowed their scope to a single strain called S288C and its derivatives. “They were domesticated for research,” said Gianni Liti, a yeast geneticist at the Institute for Research on Cancer and Aging in Nice, France. “They lost lots of the characteristics of the wild ones.”
But more recently, as Liti explains in his paper on yeast in the eLife series, natural variation has again come to the fore. In 2018, Liti led a study which sequenced 1,011 yeast strains from all over the world, including wild strains isolated from Malaysian palm nectar and Mediterranean oak trees, as well as human-associated strains from French bakeries and West African brewing vats. Among other findings, this trove of variation supported the notion that S. cerevisiae may have its origin not in Europe or Africa, but in China, where yeast genetic diversity was highest. Further, the 1,011 strains confirmed that S288C, despite its status as reference, is in important ways an outlier from its wild relatives. It has, for example, one set of chromosomes where most yeast have two.
Mark Johnston, a yeast geneticist and emeritus professor at the University of Colorado School of Medicine who was part of the team that published the first sequence of the complete S. cerevisiae genome, doesn’t use wild strains in his research. But he thinks there’s much to be learned from this work at the wild frontier, especially about evolution. “Natural selection has done the experiment for us over millions of years. And so sampling that diversity tells you a lot about that experiment,” he said. But he cautioned against expecting the kinds of fundamental discoveries about how life works that first made the model organisms famous. “I think the general principles for most of those things are worked out,” he said. “And so what we're getting there is the details.”
For Baldwin though, it’s precisely the details that count. For the first time biologists can go back to those questions that motivated 19th-century naturalists like Alexander von Humboldt and Charles Darwin, whose inquiries sent them around the world in search of the buzzing, flapping, slithering peculiarities of life on earth, he said. It’s just that now, biologists have powerful tools to investigate those questions of natural history from the chemical and genetic mechanisms within a single cell to the epic sweep of evolution. “All of the beautiful granular detail of how” different organisms deal with different environments has not been accessible with the kinds of investigation possible in the classic model organisms, said Baldwin. “I think that is just what's about to happen now.”
This article was originally published on Undark. Read the original article.

Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

