From the journals: JBC
Sphingosine might block SARS-CoV-2. Exosite binding activates factor X. How does Fur sense iron? Read about papers on these and other topics in the Journal of Biological Chemistry.
Sphingosine, spike protein and SARS-CoV-2
As global COVID-19 cases continue to surge, researchers race to identify new treatments that will expand clinicians' therapeutic arsenal. Both groups are keenly interested in developing a prophylactic treatment that can guard hosts against infection by SARS-CoV-2, the virus that causes COVID-19, or least prevent infections from taking the most severe course of disease that potentially could lead to death and disability.
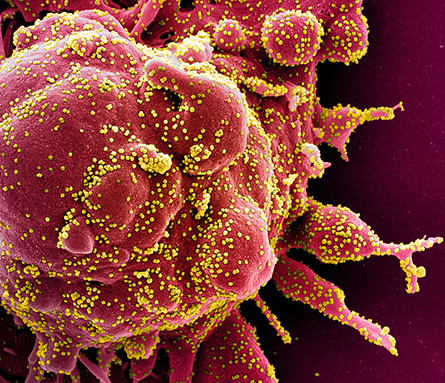
an apoptotic cell (red) heavily infected by SARS-CoV-2 virus particles
(yellow), isolated from a patient sample.
SARS-CoV-2 infects cells through an interaction of the viral spike glycoprotein with its cellular receptor angiotensin-converting enzyme 2, or ACE2. Association of these two proteins leads the virus to enter its cellular host, where it goes on to replicate. Identifying ways of interrupting this interaction could lead to new strategies for preventing SARS-CoV-2 infection.
Sphingosine, a primary component of sphingolipids, has the demonstrated ability to protect against bacterial respiratory tract infections, but researchers don't know if it also can prevent and eliminate viral infections. Michael J. Edwards of the University of Cincinnati Medical School and collaboratorsinvestigated the ability of sphingosine to modulate the infection of human epithelial cells by pseudovirus particles expressing the SARS-CoV-2 spike glycoprotein. The team showed that pretreatment of cultured epithelial cells from African green monkeys or freshly isolated nasal epithelial cells with sphingosine (suspended in 0.9% sodium chloride solution) can prevent infection by SARS-CoV-2 pseudovirus particles. They went on to identify the mechanism responsible for this result, showing that sphingosine binds to ACE2, thus blocking its interaction with the viral spike protein.
These findings, published in the Journal of Biological Chemistry, suggest that sphingosine may have therapeutic potential for prevention of infections by viruses, including SARS-CoV-2.
Sorting out the role of Serincs in HIV–host fusion
Human immunodeficiency virus reduces an infected individual's ability to ward off infections and, left untreated, can lead to AIDS. Prior to infection, HIV must enter a host cell by fusing its lipid envelope with the cell's plasma membrane. This process happens so rapidly that researchers have difficulty identifying its intermediate steps.
In a recent paper published in the Journal of Biological Chemistry, Amanda Ward of the University of Virginia School of Medicine and colleagues used cryo-electron tomography and total internal reflection fluorescence microscopy to visualize HIV–membrane fusion and unravel the role of Serinc proteins in viral infection. The researchers adapted giant plasma membrane vesicles from cell membranes that were modified to express the appropriate receptors as targets for fusion with HIV envelope glycoprotein-expressing pseudovirus particles. By controlling whether the particles included Serinc host restriction factors, the investigators showed with unprecedented clarity how Serinc proteins, specifically Serincs 3 and 5, can interrupt membrane fusion.
Exosites mark the spot for factor X activation
The interplay between active sites and exosites, secondary binding sites located far from active sites, is crucial for protease specificity. Researchers have been unable to define the balance between these sites for serine proteases that act as coagulation factors in the blood.
Manjunath Goolyam Basavaraj and Sriram Krishnaswamy of the Children's Hospital of Philadelphia found a new mechanism by which coagulation factor X, which plays a key role in the coagulation cascade, is activated. Using biochemical, pharmacological and fluorescence studies, the scientists demonstrated that this activation, mediated by an intrinsic Xase complex, requires exosite binding.
These results, published in the Journal of Biological Chemistry, may help with future inhibitor development that will aid in the treatment of coagulation disorders such as hemophilia.
Iron sensing in E. coli
The bacterial ferric uptake regulator, or Fur, transcriptionally modulates genes to regulate iron uptake, storage and use in response to changes in ferrous iron. However, researchers have not identified the mechanisms underpinning the iron-sensing ability of Fur.
Using Fur purified from E. coli, Chelsey R. Fontenot of Louisiana State University and collaborators at the University of St. Thomas showed that Fur binds an all-Cys-coordinated [2Fe-2S] cluster via conserved cysteine residues. However, depletion of intracellular free iron using a membrane-permeable iron chelator removed the [2Fe-2S] from Fur in E. coli cells.
These findings, published in the Journal of Biological Chemistry, provide new mechanistic information on how the regulator responds to changes in iron.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

