Crystal building blocks of triglycerides
Francis Crick once said, “If you want to understand function, study structure.” Do you agree? I certainly do, and I would argue that most lipid biologists do too. Just consider the effort to define chemical structures for the many thousands of lipids that exist and the hypotheses about individual lipid function that this structural information has generated.
What has lagged behind is the characterization of the structures of the proteins that modify, transport or interact with these lipids, but times are changing. For example, when I started my postdoc in Yusuf Hannun’s lab, only a handful of sphingolipid-metabolizing enzymes had been structurally characterized, and these were mainly from bacteria. While many questions remain open (hey, ceramide synthase — we can’t wait to see what you look like!), work from several labs has defined the structures and mechanisms for many human enzymes in sphingolipid metabolism.
A similar revolution appears to be happening with triglycerides. As most of you know, triglycerides serve as a reservoir for energy storage, but when they accumulate excessively, they can cause health problems, including obesity, diabetes and heart disease. Three new structures in particular have caught my attention.
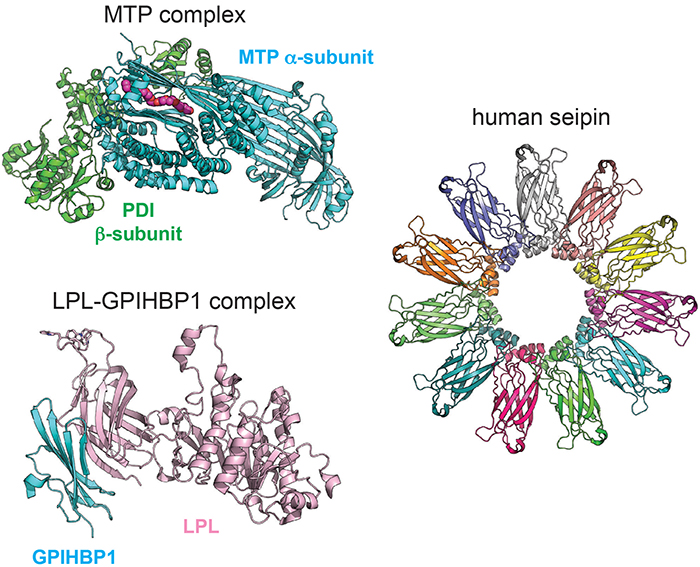 Beautiful structures of proteins involved in triglyceride metabolism and storage.Michael AirolaThe most recent is a crystal structure of microsomal triglyceride transfer protein complex, which transfers neutral lipids into apolipoprotein B-containing lipoproteins. The arduous crystallography required to conduct this work is impressive. The researchers revealed an unexpected lipid-binding cavity and provided insight into disease mutations as well as pharmacological inhibition of this therapeutic target.
Beautiful structures of proteins involved in triglyceride metabolism and storage.Michael AirolaThe most recent is a crystal structure of microsomal triglyceride transfer protein complex, which transfers neutral lipids into apolipoprotein B-containing lipoproteins. The arduous crystallography required to conduct this work is impressive. The researchers revealed an unexpected lipid-binding cavity and provided insight into disease mutations as well as pharmacological inhibition of this therapeutic target.
The second is the crystal structure of lipoprotein lipase, or LPL, the major lipase that clears triglycerides in the blood. Gabriel Birrane and colleagues and Risha Arora and colleagues separately determined the LPL structures, overcoming the relative instability of LPL by complexing it with its binding partner glycosylphosphatidylinositol-anchored high-density lipoprotein-binding protein 1. These structures, along with other biochemical data, suggest LPL is active as a monomer, challenging the long-standing paradigm that LPL was only active as a dimer.
The last notable structure is that of seipin, a homo-oligomeric integral membrane protein that is a key player in the formation of cytoplasmic lipid droplets. Two groups (Renhong Yan and colleagues and Xuewu Sui and colleagues), using cryo-electron microscopy, found that 11 or 12 seipin molecules (dependent on the species) come together to form a ring that spans the endoplasmic reticulum membrane, can bind phosphatidic acid and may stabilize the formation of nascent lipid droplets.
What’s next? Who knows, but I’m darn sure we’re all gonna love it.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

