Move over, DNA. The future is protein
In the coming decades, efforts to understand disease will be propelled by building 3D maps of protein arrangements in cells over time, a team of scientists says.
A workshop about the study of proteins on a large scale — proteomics — yielded a perspective piece that is part vision, part plan, recently published in the journal Molecular & Cellular Proteomics.
DNA is like a game plan, explains Neil Kelleher, a researcher at Northwestern University who was involved in the study, and RNA is like the team huddle; it’s what the cell decides to do with the circumstances on the day.
It’s time to watch the game itself. “This is the decade,” Kelleher said.
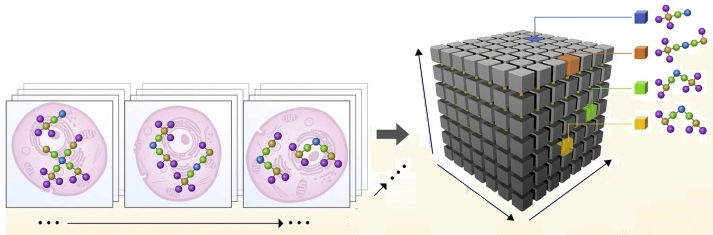
Watching the game
“Traditionally, we study one protein or pathway at a time,” the authors write, because “the cell is otherwise too complex.”
That’s like watching one player over time and whom they pass to.
Proteins take multiple forms in a cell after they come off the bench. An electric link between fans and footballers inspires the play that decides a championship. A comment or word from a coach flicks a switch. In the cell, such switches can be as simple as a few atoms added (phosphorus and a trio of oxygens are a popular one). Altered by their circumstances, proteins are like people.
Mapping their shapes over time will be game-changing.
“This could impact all diseases,” said Kristin Burnum–Johnson of Pacific Northwest National Laboratory, first author of the paper.How will this happen?
Bottoms up, touchdowns
Scientists will tackle this from two directions: bottom-up and top-down. They start with clinical tissue samples, Burnum–Johnson explained.
"For bottom-up approaches, you extract proteins from tissue regions and break them into identifiable peptides (shorter strings of proteins).”
Bottom-up strategies, combined with RNA and metabolite data, will reveal the lineup. Top-down strategies will yield those sweet, sweet action shots.
Kelleher knows about this. The top-down approach, revealing the specific forms proteins take (which RNA sequencing can’t tell you), has become so sensitive that his research recently found 30,000 precise forms of proteins in blood and bone marrow alone. That’s almost 10 times more than in previous studies. There are thousands more cell types for postgame analysis.
Single-minded
Researchers will continue pushing for higher resolution. They’ll need to integrate data, which is not a simple challenge. They’ll need to think beyond stadium walls; the extracellular matrix is rich. Like football and cells alike, consortiums and partnerships will drive the process. The National Institutes of Health wants research gaps to be identified, and private investment is supporting the sequencing of single molecules. These combined efforts remind Kelleher of the Human Genome Project.
The central dogma of biology is DNA to RNA to protein. It’s only natural that scientific research follows the path that is familiar. It’s protein’s turn.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

