
Personal chemistry
Philipp Geyer was interested in learning how protein levels in the blood change when a person tries to lose weight. Several years ago, Geyer, then a postdoctoral fellow, and his colleagues in Matthias Mann’s lab at the Max Planck Institute of Biochemistry began to analyze samples from a dieting study, hoping to identify biomarkers that could predict the outcome of dieting and other interventions. They found something a little different from what they were looking for.
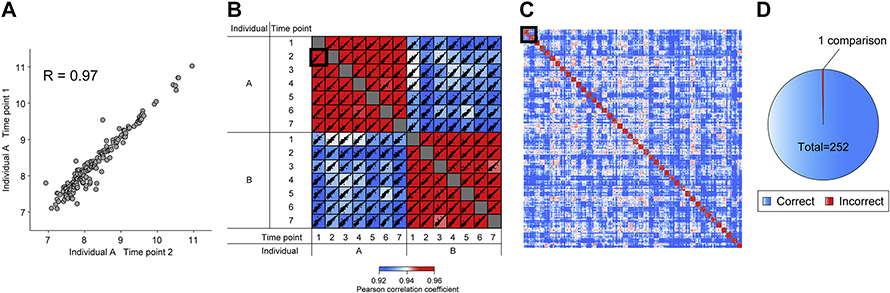
While studying the plasma of 1,500 dieters tracked for 14 months, Geyer and his colleagues observed much more variation in protein levels across individuals than within any one person over time. Although some protein levels changed dramatically in response to dietary intervention, many others remained steady. “For example, alpha-two microglobin,” Geyer said. “It’s tenfold different between people, but completely stable within a person.”
As patterns emerged from the data, allowing them to pick out the same person at different time points, Geyer and his colleagues began to worry: Could these patterns one day be used to de-anonymize a study participant?
Around the same time, leaders of an international consortium of proteomics data repositories called ProteomeXchange were convening in Amsterdam to discuss ethical and legal issues in handling potentially identifiable biochemistry data.
The meeting’s organizer, Juan Antonio Vizcaíno, runs a data repository from his lab at the European Molecular Biology Laboratory–European Bioinformatics Institute, known as EMBL-EBI, in the United Kingdom. Re-identification is “an issue that we have been aware of for years,” he said, “but there needs to be a critical mass to start discussing it properly.”
Sharing data is an important norm for the proteomics community. The benefits of openness are paid out in reproducibility, quality control and maximal use of each data set. Limiting access to data could impede scientific discoveries and their resulting therapies or even cures.
Still, even the most fervent defenders of open sharing agree that a communitywide discussion about access to proteomics data is worth having.
Earlier this year, the journal Molecular & Cellular Proteomics, which has played a significant role in creating a culture of openness, published three articles — two from the Mann lab and one from Vizcaíno and colleagues — on the topic.
ASBMB Today talked to the authors of those papers, the editor who oversaw them, legal and ethics experts, and other researchers about what is and isn’t technically possible today, the risks and rewards of open proteomics data, and how to make scientific progress while still protecting people’s privacy.
What can a proteome reveal?
Whereas genome sequences are widely considered recognizable and linkable to an individual, most researchers so far have considered proteomes more anonymous.
Robert Gerszten, a physician–scientist at the Beth Israel Deaconess Medical Center, compared matching the pattern of protein levels in a clinical proteomics experiment to identifying a blurred photo. If a genome is like an image of an individual’s face, a proteome, because of biological and technical factors, is more like a partially masked image at lower resolution. It takes longer and costs more to collect a proteome than a genome, and the depth of coverage — the number of times an experiment confirms that a specific gene or protein is present — is much lower in mass spectrometry–based proteomics. Due to technical factors in protein preparation and measurement, not all proteins are equally likely to appear on a spectrum, and absence from the data doesn’t mean that a protein is not present in the sample.
“If you take a population of thousands of people,” Gerszten said, “you can clearly find person-specific signatures in that population (based on protein concentration). You couldn’t have done that five years ago. … But I don’t think that the granularity is enough to figure out who that individual might have been in a huge population.”
Stefani Thomas, whose lab at the University of Minnesota uses proteomics to investigate candidate biomarkers for cancer diagnosis and prognosis, said the Mann lab’s diet study offers a thought-provoking proof of concept that protein level alone might be identifiable. However, she said, to determine traits such as an individual’s age, gender or ethnicity using their proteome, researchers would need to know more about variability between and within those groups — similar to the research her lab does to differentiate between healthy and disease states.
Researchers don’t know exactly how stable a level-based proteomic signature is over time. The Mann lab’s study followed participants for over a year, which is brief compared to a lifetime. And though weight loss left most parts of the proteomic fingerprint unchanged, the impacts of other physiological events are not yet established.
In addition to a single protein’s presence, abundance and post-translational modification status, proteomic experiments increasingly can reveal something that doesn’t change over a lifetime: genetic sequences, in the form of genetically variable peptides.
Proteomics yields sparser sequence information than DNA sequencing. The exome is smaller and subject to more selective pressure than the genome, and protein translation makes synonymous mutations invisible, so less person-to-person variation exists in proteins than in DNA or RNA. Still, when Geyer and colleagues went back over their data looking for genetic variants, they could indeed identify individual single-amino-acid variants in some spectra.
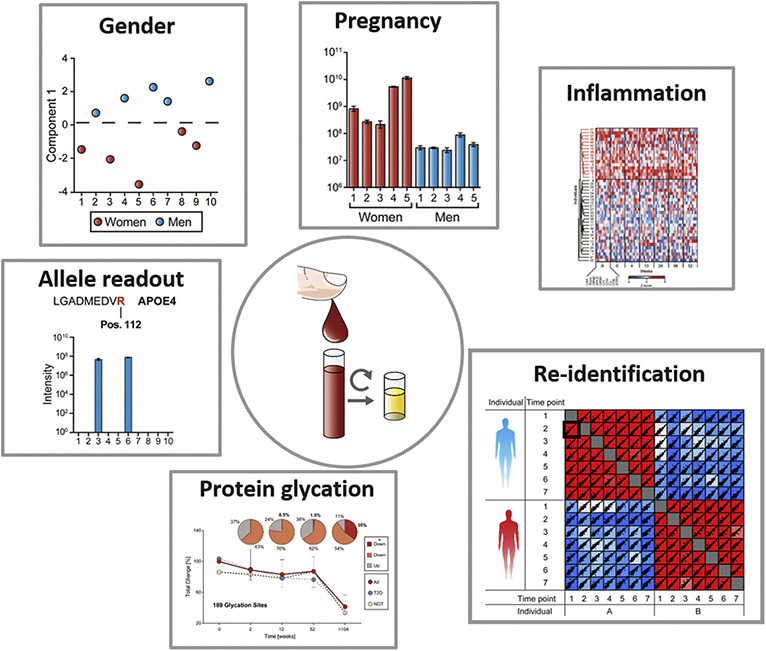
studies to determine potentially identifying features such as gender, pregnancy status and allele expression in a cohort.
Geyer got to talking with Sebastian Porsdam Mann, Matthias Mann’s son, who recently had finished a Ph.D. in bioethics, about how much of a risk this posed to participants — and whether the possibility of discovery outweighed the risk.
“You need to have high quality data, but in order to reduce identifiability risk, you need to do things to this data to filter out identifiable information. So there’s always going to be a trade-off,” Porsdam Mann said. In addition, there are times when being able to match a study participant to features of their proteome could be beneficial.
Treatable discoveries
Sometimes researchers accidentally make an observation that could affect a study participant’s health. Philipp Geyer made such a discovery — but the participant was himself.
Read moreIn the Mann lab, as in other labs handling clinical samples, specimens arrive stripped of participant name, date of birth and other identifying information. Do the specimens themselves — or, more importantly, the data the lab uploads to international repositories — carry enough information that a motivated, proteomics-savvy adversary could trace them back from spectrum to person?
According to Thomas, a key question remains unresolved: “What is the minimum amount of data from a proteomic experiment that can be used to definitively identify an individual?”
Although the clinical proteomics community is just beginning to explore the question, forensic researchers have been trying to determine an answer for years.
Forensic proteomics
Glendon Parker, an adjunct associate professor at the University of California, Davis, is one of a small number of scientists working to identify genetically variable peptides for forensic purposes; his lab focuses on hair. Inferring a genome, Parker said, depends on knowing whether each change to a peptide is genetic or is caused by environmental effects such as dyes or weathering. That means validating that each candidate genetically variable peptide matches a genotype before using it to infer genetic information — and also that it is distinguishable from the rest of the proteome.
Deon Anex, a chemist at Lawrence Livermore National Laboratory who works on a forensic proteomics project inspired by Parker’s research, frames the question: “Let’s say you have a unique peptide, and then you modify one of the amino acids. Is that still unique? Or is it now a common sequence somewhere else?”
In most cases, using the proteome to get a genotype is redundant to long-established methods based on DNA. However, in some environments, DNA falls to pieces but proteins survive. According to Anex, forensic proteomics is a good fallback for getting genetic information from hair or from brass ammunition cartridges that are inhospitable to DNA. His lab also is working on a project funded by the research agency of the Office of the U.S. Director of National Intelligence that aims to pull genetic information from the proteins left after bomb blasts. The project team has demonstrated that it can match proteins from partial fingerprints on objects to the volunteers who handled the objects. Now, it’s conducting field experiments with lab-made improvised explosive devices to determine whether those traces are still recognizable after an explosion.
Based on the variant peptides they detect in a sample, forensic researchers in Parker’s and Anex’s labs can determine a partial genotype. Using statistical methods from forensic science, which consider the number of variants detected and how common each one is, researchers can calculate the odds of a false-positive match. Those odds would need to be on the order of one in 10 billion to identify someone uniquely in the world. Parker has an answer to the question Thomas posed; he believes that just a few hundred genetically variable peptide calls could be enough to identify some people uniquely.
But even when the odds of a false positive are very low — even with a genome that unambiguously belongs to a single person — it is difficult to trace a sequence back to the person who carries it in their body. Brad Malin is a professor of biomedical informatics at Vanderbilt University. “Uniqueness is insufficient to actually identify somebody,” he said. “You need to be able to link that data to some other resource to get back to their identity.”
He added later, “There have been illustrations that when an organization or an individual is sufficiently motivated, if all the stars align, they’ll be successful. But it requires a lot of effort.”
Back in the clinical proteomics community, some researchers say that the risk of a motivated, proteomics-savvy adversary making such an effort is dwarfed by the benefits of open data sharing. Some also ask whether, in a world where so much personal information already is collected and sold, the risks of adding proteomics data to the mix could possibly outweigh the advances in science and health that those data enable.
What’s the harm?
Malin said a friend recently questioned him about why re-identifiability was a concern, asking, “What are the harms? What is it that we’re supposed to worry will go bump in the night?”
For one proteomics researcher, Michael Snyder, the question is personal. Snyder, a Stanford University professor, conducts multiomics studies that include himself as a study subject; he published his whole genome, thinly anonymized, years ago in the journal Cell and has followed it up with longitudinal transcriptomes, metabolomes and clinical assays.
Sporadically over the years, Snyder has heard from people who want to share analyses of his health. Some have had interesting insights, he said, but “some of it’s pretty loony.” Still, he doesn’t believe he has suffered any real harm from radical openness about his personal biochemistry. Nor is he aware of other study subjects who have suffered because their transcriptomic or proteomic data have been published.
“There’s a lot of RNA-Seq data out there,” he said. “I challenge you to show me one example where that’s been abused.”
He’s right; no examples of such abuse have been reported. The loudest bumps in the night come from privacy researchers sounding the alarm about data vulnerability and law enforcement agencies using genetic information from opt-in genealogy databases — not research data. Other possible attackers are so far a matter of conjecture.

or PRIDE, out of the European Molecular Biology Laboratory in Cambridge, UK.
Information in a proteome can give clues about a person’s health; American scholars have envisioned discrimination by health insurers as a concern, but it is now illegal for insurers to deny coverage based on genetic information. Gerszten, the physician–scientist at Beth Israel, suggested that marketers hypothetically could mine metabolomic or proteomic databases to find out about reproductive choices or disease status. There is no doubt that a market for this type of data exists; according to a cybersecurity firm’s 2019 report, health records stolen from hospitals sell for about 50 times the value of a stolen credit card number. (Of course, health records are easier to interpret than a proteome, whose significance may not be known, and they generally include a patient’s name or other identifiers.)
After revelations that social media and genealogy sites can gather and reveal more information than users thought they were sharing, societal conversations around privacy and data sharing have changed in the past decade. Snyder has noticed a similar uptick in caution among his study participants since he launched a longitudinal proteomic profiling project called iPOP in 2010.
“It did take this shift when people learned that Facebook and others were selling their data,” Snyder said. However, he added that he sees participants on social media “posting some incredibly private stuff that’s probably more harmful than proteomics data. Privacy is gone, whether you like it or not.”

Malin, the bioinformatics expert, disagreed. “Google’s not taking all of your search queries and throwing them online for everybody to see,” he said. “Privacy is not dead. You’ve just shifted who you trust with information about yourself.”
From an ethical standpoint, Malin said, any research participant whose privacy is compromised has lost something of value, even if they suffer no further consequences. “Whether or not the individual was materially harmed ... simply the identification would be sufficient to claim that their privacy had been infringed upon, because they did not want that information revealed.”
Lawsuits against hacked hospitals have argued the same thing. But researchers say that they, too, have something of value to lose: knowledge that could lead to biomedical progress. Broad Institute proteomics director Steven Carr, an MCP deputy editor, said, “At this point in time, adding unnecessary protections to the availability and use of proteomics data on human samples has the potential to do more harm than good.”
What do researchers stand to lose?
In the past year and a half, numerous large COVID-19 studies have illustrated the benefits of data sharing in proteomics. Data sets showing viral interaction with human proteins, linking detectable markers to patient outcomes and tracking immune responses over time have added to an internationally constructed picture of how the novel coronavirus works and have been mined by other researchers for further insights. And this is not a new phenomenon; according to Snyder, most of the annotation of the human proteome has depended on secondary analysis of publicly available data.

Since its launch in 2011, the ProteomeXchange, a consortium of repositories that make proteomic data freely available, has published more than 24,000 mass spectrometric data sets, about 45% of those from humans or human-derived cell lines. Thousands of papers have reported new findings based on data in the six repositories that make up the exchange.
The field was not always so open. It took commitment from publishers, funders, data repositories and the Human Proteome Organization to require investigators to make their raw data freely available to colleagues. The editors of MCP, Carr said, take pride in the journal's early requirement that scientists publish raw data.
“There was a point where very few people submitted data to PRIDE or to other resources,” Vizcaíno said. After a rapid change, “Now we are in a scenario that is basically the opposite.”
Carr and Snyder argue that limits on data sharing to accommodate privacy concerns would risk reversing the field’s cultural shift. Without easy access to raw data, researchers would be unable to check one another’s work for reproducibility or reanalyze data for follow-up studies. Based on how usage patterns differ between controlled-access and openly available transcriptomics data, Snyder is confident that research progress would slow if proteomics data were walled off.
Some proteomics data already are controlled because they are linked to other, more easily identifiable data such as clinical outcomes. Gerszten, whose lab conducts multiomics studies of heart disease, said that they keep all data from patients under what he called electronic lock and key. “The ability to say, ‘these sets of markers track with individuals who had a better outcome’ … is actually very valuable information,” Carr said. “It represents no harm to any individual. But it does represent a pathway to try to identify markers that might be useful for diagnostic purposes.”
Gilbert Omenn, a physician–scientist who directs the University of Michigan’s center for computational medicine, said, “We want proteomic data not to just be about advancing tools of mass spectrometry and other methods; we want it to be linked to biomarker development and clinical diagnosis, and therefore we need to translate it to patients.”
That goal is coming into view, he said. It is exciting for the field, but “I don’t think it behooves us to try to claim that we’re outside the boundaries of responsibility.”
Malin believes that a major re-identification episode would do more harm to researchers than to their participants. “People are going to scream bloody murder that they will no longer trust scientists with their information,” he predicted. “That would be a big blow to biomedical research.”
Legal requirements, technological solutions
Laws governing data sharing
HIPAA, GDPR and the Common Rule all affect how scientists in different regions are required to protect, and allowed to share, potentially identifiable data.
Read moreConcerns about potential damage to the field motivated Vizcaíno to organize the Amsterdam meeting of ProteomeXchange leaders and other experts in 2019.
In a recent article in MCP inspired by that meeting, the researchers called for data access to become “as open as possible, as closed as necessary” to balance research transparency and data sharing with privacy concerns. The field needs further research into the likelihood and potential severity of data breaches, they concluded, but even in the absence of such research, it is important to begin to develop best practices for handling sensitive data.
Proteomics databases are designed to make data public and permanent. After a quality control check, PRIDE publishes all the data it receives, sometimes waiting for a corresponding paper to be published. Here and there over the years, Vizcaíno said, “We have had a few cases that people who had originally submitted data to us have told us, ‘We have been told by our data officer … that we shouldn’t do this.’”
University data officers are trying to follow complex rules. In many jurisdictions, consumer privacy laws address what identifiable data may be shared (see “Laws governing data sharing”), and different authorities balance the risks and rewards of data sharing differently.
Omenn said there is “a lot of attention, a fair amount of angst, and, I think, rather high compliance” with legal requirements for identity protection.
Because policy changes slowly and tends to be more reactive than proactive in the face of technological advances, Malin expects that it would take a major breach of privacy affecting either millions of people or a powerful politician to force changes to privacy laws. When an American policy called the Common Rule was revised, a process that took six years, the agencies involved in the work decided against declaring genetic information identifiable.
“The implications of designating biological information as identifiable are quite breathtaking,” Malin said. “It would completely shift the way that research is performed in this country.”
Still, he said, “We’re somewhat at a crossroads … it is possible that the United States is going to have its hand forced by the European Union.”

Vizcaíno, predicting that re-identification would become a concern for proteomics, has kept an eye on the DNA and RNA databases that his EMBL-EBI colleagues administer. Often, these administrators require that researchers apply for access to sensitive data, providing only enough information to run analyses the labs describe ahead of time. Some databases layer in additional measures, such as suppression of highly identifiable sequences or scrambling of genotypes, to protect information that could identify an individual; meanwhile, in a sort of arms race, privacy researchers continue to report ways to breach those safeguards.
For now, PRIDE and other databases do not have the architecture in place to render raw proteomics data less identifiable. If Vizcaíno and his colleagues receive a request to delete a data set while it’s still under review by editors and reviewers, they are able to honor that request.
Removing data sets after they are posted, however, may raise problems with the resulting publications. Carr said, “MCP will not accept papers where the data cannot be made public.”
Vizcaíno hopes to build a database that, like databases for genomics, offers controlled access only to reviewers and researchers who explain why they need to see sensitive data; researchers might be restricted to accessing data that corresponds directly to a research question. Approaches from genetics, he said, are practical, but an investment is needed to adapt them to mass spectrometry data formats. “We are at the very beginning of a very long road.”
Sightlines
Ongoing conversations about proteomic privacy are driven by what biomarker researcher Stefani Thomas called “an undercurrent of advances in technology” and a faster pace of proteomic data collection. That’s good news for the field, she said, but to realize the goal of using the technology in the clinic, questions about privacy and identifiability must be resolved. “I think it’s exciting that people in the field are taking a step back and saying, ‘Let’s look at this from a broader perspective and make sure that what we’re doing is ethical.’”
Porsdam Mann, the bioethicist, said, “If you look at the history of genomics research, but also other related fields, you’ll find that responsible self-regulation early in the game is one of the wisest longer-term investments.”
Carr said, “The way I view this is in clinical terms. … I think we should be in a position of watchful waiting to see how confidence in potential identification of proteomics data goes.”
Snyder argues that existing restrictions on data use are more than adequate. “Until somebody gets harmed, I’m not sure I’m so worried about it,” he said. “Maybe when somebody gets harmed, it will all blow up and they’ll say, ‘Mike, you didn’t foresee this very well.’ And I’ll say, ‘Yeah — but we got a hell of a lot done in the meantime.’”
Laws and policies governing data sharing

The Bermuda Principles, established in 1996, supported rapid, public data sharing to advance the Human Genome Project. In the years since, concerns about credit for research and about participant identifiability have driven a shift to more cautious data sharing.
The European Union’s General Data Protection Regulation took effect in 2018. Its protections on data collected in Europe and describing European citizens make exceptions for anonymized research data, but experts say that anonymization is not well defined in the law, making it hard to follow. In a Policy Forum article in the journal Science, legal scholars in the U.S. and Europe argued that GDPR protections impeded collaborative international research by preventing clinical data collected in Europe from being transferred to international collaborators. The barriers to transferring data, they wrote, “appear to be at odds with the generally research-friendly intent of the GDPR.” According to Vanderbilt bioinformatics professor Brad Malin, until a court rules on anonymization requirements, the exact meaning of the GDPR for genetic and bioinformatics data will remain ambiguous.
In the United States, data sharing is governed by a federal policy, originally adopted in 1991 and since revised, known as the Common Rule. The policy, which applies to any research on human subjects carried out or funded by 15 federal departments, is forged by consensus among them. The rule exempts data “recorded such that subjects cannot be identified.” As with the GDPR, the Common Rule is complex and does not spell out whether genomes are identifiable or can be anonymized. Health records, such as those produced in clinical trials, also are protected by the Health Insurance Portability and Accountability Act of 1996, or HIPAA; it, too, makes provision for sharing of de-identified data for research purposes.
Treatable discoveries
Philipp Geyer, a researcher at the Technische Universität Berlin, had high cholesterol starting in childhood. As a postdoc, while setting up a workflow for a large clinical trial, he used a sample of his own blood as a quality control. By revealing the amino acid sequence of a peptide from apolipoprotein E, the proteomic data showed that Geyer had a variant of the gene linked to persistent high cholesterol levels.
After a lifetime of limiting chocolate and butter, pursuing sport and exercise regimens, and stubbornly declining pharmacological intervention, Geyer said, the discovery that his problem was genetic gave him a push. “I was convinced, okay, I can’t do anything with sports or nutrition, so I really have to take the drug.”
On a statin, Geyer’s cholesterol problem resolved. Monitoring his own plasma proteome, he watched his apolipoproteins drop. He shared this information with members of his family. “My dad and my brother actually went on statins because of this,” he said.
Geyer’s discovery is what an ethicist would describe as an incidental finding. It wasn’t the information he set out to find, but it gave an important insight into his health. And a simple intervention helped Geyer lower his risk of cardiovascular disease.
According to Sebastian Porsdam Mann, a bioethicist and son of PI Matthias Mann who worked with Geyer on a Molecular & Cellular Proteomics article about the ethics of clinical proteomics studies, this is part of why absolute anonymization of data may not always be in a study participant’s best interest. “In the extreme, you could literally anonymize it completely,” he said. “But then you could never follow up.”
Steven Carr, proteomics director at the Broad Institute, has experienced the frustration of being unable to follow up; while analyzing a lung cancer sample, he said, his lab stumbled across a clinically meaningful finding. He said that his team wanted to get in touch with the clinicians who collected the sample to tell them, “We don’t know who this person is, but by the way, they could have been — or should have been — treated with X because they have this particular set of characteristics in their proteome.” Because of privacy protections, he said, that kind of feedback was prohibited.
Some incidental findings are less actionable. For example, people who carry a different apolipoprotein E allele than the one Geyer discovered in his blood are up to 90% more likely to develop Alzheimer’s disease, but the connection is not well understood and someone who learns they have the allele can’t take any action but wait.
Medical ethicists see a line between actionable and nonactionable information; they say that in general, if information is actionable, patients ought to be informed — unless, fulfilling the principle of autonomy, they have stated they prefer not to be informed.
Certain incidental findings might convey information a study participant would prefer not to share with others. For example, physician–scientist Robert Gerszten mentioned that metabolomics studies designed to find heart disease biomarkers also might show what medications a person is taking or whether metabolites of illicit drugs are present.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.


