From the journals: MCP
A triphosphorylated species may be linked to DNA repair. A site-specific proteomics approach holds promise for Alzheimer’s diagnosis. A new platform improves quantification of histone modifications. Read about papers on these topics recently published in Molecular & Cellular Proteomics.
A lipid in the nucleus
Phosphatidylinositol is a complex molecule. The lipid’s head group, a sugar, has three potential phosphorylation sites, generating seven potential phosphorylated species called polyphosphoinositides, each of which is thought to bind a different set of effector proteins and exert diverse effects.
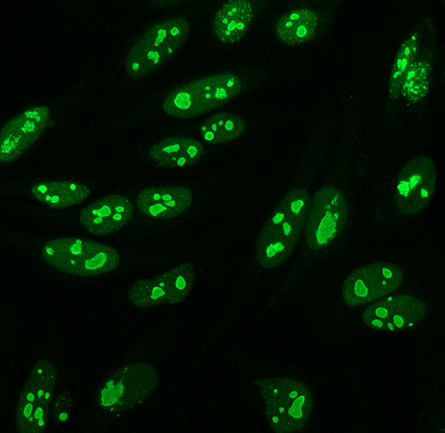
Researchers long have known that cleavage of the head group from its hydrophobic fatty acid tails forms a cytoplasmic second messenger. More recent research has identified polyphosphoinositides, complete with fatty acids, in nonmembrane environments. Researchers have mapped the interactions of a few phosphoinositide species with cytoplasmic and nuclear proteins, but they do not understand yet the role of the triphosphorylated species PIP3.
In a recent article in the journal Molecular & Cellular Proteomics, postdoctoral researcher Fatemeh Mazloumi Gavgani and colleagues at the University of Bergen in Norway describe how they used immunofluorescence to determine that nuclear PIP3 mostly is located in the nucleolus, where ribosomes are assembled. Then they used mass spectrometry to identify PIP3-interacting proteins from the nuclear fraction of HeLa cells; most of the proteins were nucleolus residents with known roles in RNA processing, splicing or catabolism, or translation.
One binding partner was poly(ADP-ribose) polymerase 1, or PARP1, an enzyme that adds a chain of ADP-ribosyl subunits to protein substrates after DNA damage, mobilizing proteins involved in damage repair. Through truncation analysis and directed mutations, the authors found that three polybasic regions in PARP1 rich in arginine and lysine residues mediate its binding to PIP3. The authors argue that the interaction suggests a link between PIP3 and DNA repair that merits further study.
Given that the bulk of each PIP3 molecule is hydrophobic, the researchers still aren’t sure how it might be transported to or synthesized in the nucleolus and remain in solution. They speculate that the lipid could form micelles or be shielded by binding to carrier proteins.
New approach holds promise for Alzheimer’s diagnosis
The cerebrospinal fluid, or CSF, that surrounds and supports the central nervous system, or CNS, plays an important role in brain development and neuronal functioning. It is the only body fluid that directly interchanges with the extracellular fluid of the CNS and reflects ongoing pathological changes in the CNS. Thus, proteomic analysis of CSF has great potential for CNS-related disease diagnostics.
Glycosylation, an important post-translational modification, regulates many cellular processes. Defects in glycosylation have been linked to human diseases, making it a valuable target to study. While researchers have done extensive proteome profiling for CSF, few studies aim at unraveling a site-specific CSF N-glycoproteome.
In their recent paper in Molecular & Cellular Proteomics, Zhengwei Chen, Qinying Yu and a team at the University of Wisconsin–Madison describe their large-scale site-specific approach for in-depth CSF N-glycoproteome analysis, which allows for thorough coverage of CSF N-glycopeptides, including glycopeptide sequences, glycosylation site and glycan composition. In addition to generating the largest reported N-glycoproteome data set for CSF to date, the researchers applied this strategy to compare the N-glycoproteome coverage between people with and without Alzheimer’s disease. They uncovered diverse and distinct glycosylation patterns, paving the way for promising glycosylation-based biomarker candidates for Alzheimer’s. This work lays a foundation for more in-depth investigation of the functional roles of these glycosylated proteins in the progression of Alzheimer’s and other neurodegenerative diseases.
New platform to quantify histone methylation
Histone modifications are post-translational modifications to histone proteins that affect gene expression by altering chromatin structure and are responsible for packaging long DNA molecules into more compact, denser structures. Methylation is a common histone modification mediated by the addition of one or more methyl groups on the lysine or arginine amino acid of the histone protein.
Methylation is important in gene regulation and has been linked to progression of many cancers; however, researchers have difficulty quantifying this histone modification because of the spatial complexities that result when more than one methyl group is added on the same amino acid.
Francesca Zappacosta and a team of researchers at GlaxoSmithKline recently developed a simplified mass spectrometry–based platform for histone methylation analysis. The technique uses chemical derivatization to reduce the complexity of the protein to be analyzed, improving sensitivity for the detection of methylation. In one tested approach, the researchers quantified and distinguished between symmetrical and asymmetrical dimethylation of histone H4R3. They also were able to quantify modification levels as low as 0.02%. The team describes their mass spectrometric platform and findings in a recent article in Molecular & Cellular Proteomics.
This effective and robust MS-based approach holds promise for many studies that involve quantification of histone methylation profiles.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.


