Progress in identifying lipid domains (rafts) in living cells
Under which conditions lipid chemical heterogeneity results in the formation of coexisting lipid domains with distinct lipid compositions and properties in living cells has been a subject of intense research for decades.
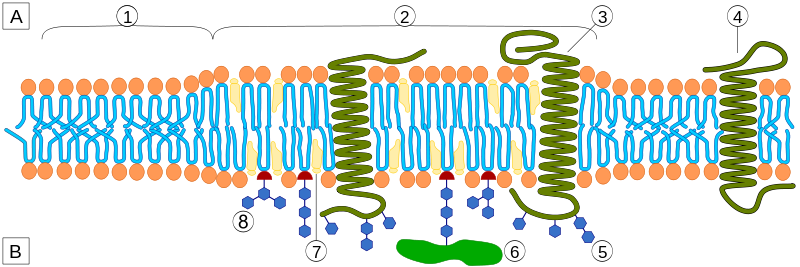
In model membrane formed from lipid mixtures, spontaneous formation of tightly packed sphingolipid- and cholesterol-rich lipid domains (in the liquid-ordered state) that segregate from loosely packed domains richer in unsaturated phospholipids (in the liquid-disordered state) are detected and characterized easily.
However, analogous domains in cells are very small under most conditions — at or beyond the limit of detection for most techniques. This has led to much controversy as well as much work aiming to develop new methods to identify and characterize tiny nanodomains.
Very recent progress in living cells has been encouraging on several fronts. Studies using novel fluorescently labeled lipids with affinities for liquid-ordered domains similar to those of unlabeled lipids have revealed that specific association of raft-loving lipids with raft-localizing proteins occurs in living cells (1,2). Single-particle-tracking measurements show that these interactions are lost in living cells when even minor changes in lipid or protein structure are made if these changes abolish raft-associating physical properties.
In other studies, super-resolution microscopy in B cells has found co-localization of raft markers with, and exclusion of nonraft markers from, the vicinity of clustered B-cell receptors on a size scale similar to that of the clusters (50 nanometers to 100 nanometers). This is indicative of the formation of ordered domains around the B-cell receptors. An analogous formation of nanodomains was detected around clustered cholera toxin, a molecule long known to induce the formation of ordered domains in vitro and in cells (3).
These studies extend previous work from other labs that reported lipid-domain-based molecular interactions in these systems. This is indicative of a robust underlying phenomenon.
Advances leading to an increased ability to visualize domains and manipulate their structure promise further progress. An even higher-resolution, super-resolution microscopy approach has been developed, which may allow visualization of domains that otherwise would elude direct visualization (4).
Finally, our own lab has devised a method efficiently to replace virtually the entire complement of plasma membrane outer leaflet lipids in living cells with exogenous lipids. This may allow fine-tuned control of domain formation and properties (5).
REFERENCES
1. Komura, N. et al. Nat. Chem. Biol. 12, 402 – 410 (2016).2. Kinoshita, M. et al. J. Cell. Biol. 216, 1183 – 1204 (2017).
3. Stone, M.B. et al. eLife 6, e19891 (2017).
4. Balzarotti, F. et al. Science 355, 606 – 612 (2017).
5. Li, G. et al. Proc. Natl. Acad. Sci. USA 113, 14025 – 14030 (2016).
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

