MCP: Charting the mitochondrial interactome
As the powerhouses of the cell, mitochondria host various supramolecular protein complexes. Delineating the structural basis of these protein complexes is essential to improve our understanding of how mitochondria function and generate energy. In a study published in Molecular & Cellular Proteomics, Albert Heck and a team of investigators at Utrecht University in the Netherlands, in collaboration with the National Institutes of Health, aimed to discover the organization and interactions of proteins in the mitochondria of mouse hearts.
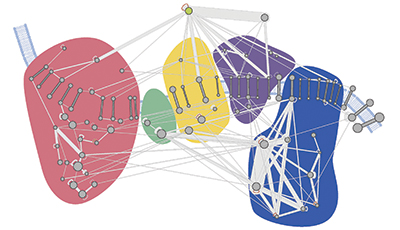 A molecular map of the oxidative phosphorylation supercomplexes. Courtesy of Philip Lössl/Utrecht University
A molecular map of the oxidative phosphorylation supercomplexes. Courtesy of Philip Lössl/Utrecht University
“We were most curious about the organization of protein molecules within mitochondria, because proteins are the molecular building blocks that make the mitochondrial energy factory work,” Heck said. “It was already known which proteins are involved in energy generation, but it is still not fully understood how these building blocks come together within intact mitochondria.”
To chart the organization of proteins within mitochondria, the researchers used a kind of molecular glue, or cross-linker, small enough to enter intact mitochondria and form stable links between any proteins within close proximity of one another. The mitochondria then were broken apart, and the proteins were digested and run on a mass spectrometer to identify especially the cross-linked peptides.
The researchers cataloged the largest set of mitochondrial protein interactions thus far, with 3,322 unique cross-links. This unprecedented depth was achieved using optimized mass spectrometry fragmentation schemes and data analysis strategies. “In contrast to earlier work based on similar strategies, our approach is much more sensitive, allowing us to present a more complete molecular interaction map of all proteins within mitochondria,” Heck said.
This molecular interaction map, or interactome, revealed a dense and interconnected network of proteins. The researchers used the map to study the higher-order organization of proteins and the architecture of protein complexes in mitochondria. Among these are the oxidative phosphorylation supercomplexes, a series of five protein complexes cumulatively responsible for generating energy. In addition to confirming known interactions, the researchers found novel cross-links between individual complexes, leading them to suggest that all five complexes coexist in close proximity.
Going a step further to validate their map, the investigators soaked the mitochondria in a high-salt solution to disrupt the protein supercomplexes. They showed, using the same cross-linking technique, that these “dysfunctional” mitochondria displayed a very different protein interaction network. “These data show that protein organization and mitochondria function are two sides of the same coin,” said co-author Philip Lössl. “We believe that our protein maps will help us understand the organization principles that allow mitochondria to work as molecular powerhouses.”
Chemical cross-linking and mass spectrometry allowed the researchers to probe native architecture of protein assemblies in mitochondria that are still intact and functioning. Most traditional biochemical methods for studying protein–protein interactions involve solubilizing the membrane using a detergent, which can introduce artifacts. “In such studies, the forceful breaking of the mitochondria can have dramatic effects on the protein organization and important information may be lost,” Lössl said. With the extensive comparative analyses and structural validation performed in their study, the researchers believe that the supercomplex interactions detected in intact mitochondria should be considered genuine.
The researchers believe their approach can be used to compare mitochondrial organization in diseases related to mitochondrial dysfunction, such as Parkinson’s and autism spectrum disorders. “Our approach can be used to elucidate how the molecular landscape of mitochondria is reprogrammed during disease development,” Heck said, “ultimately providing targets for future therapies.”
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

