From the journals: MCP
Are proteomics helping to detect ovarian cancer? The challenges and promises of biomarkers. New technique helps detect protease cleavage. Read about papers on these topics recently published in the journal Molecular & Cellular Proteomics.
Are proteomics helping to detect ovarian cancer?
Epithelial ovarian cancers, or EOCs, make up about 90% of all ovarian cancers. EOC originates in the surface layer covering the ovary and then spreads. When diagnosed early, EOC has about a 93% five-year survival rate. Early diagnosis is rare; however, due to a lack of specific symptoms. Many patients are not diagnosed until the cancer has significantly progressed, which decreases the five-year survival rate to 50%.
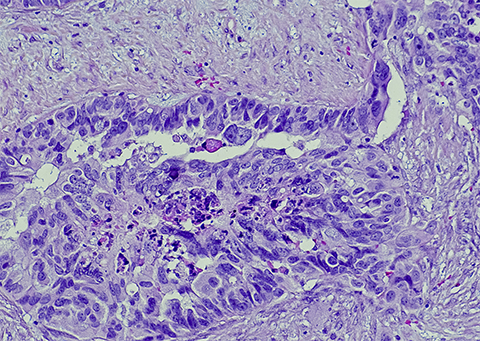
The five histological subtypes of EOC are low-grade serous ovarian carcinoma, mucinous carcinoma, endometrioid carcinoma, clear cell carcinoma and high-grade serous ovarian cancer. This last is the most aggressive and prevalent form, making up 75% of diagnosed ovarian cancers. Each subtype has different molecular underpinnings and all have varying treatment success rates. Researchers seek to identify biomarkers to facilitate early EOC diagnosis and enhance treatment options.
In a recent article published in the journal Molecular & Cellular Proteomics, Liujia Qian and colleagues from Westlake University in China performed a systematic review of over 2,500 ovarian cancer clinical trials to assess progress in the discovery of EOC biomarkers and therapeutic targets as well as the translation of these findings to clinical use.
EOC proteomics research relies on detecting biomarkers in serum and plasma where they are easily obtained, the authors explain. Researchers then use algorithms to predict the likelihood of ovarian cancer or its progression in patients based on the presence of the biomarkers. Some of these tests are already being used in the clinic, but others remain under-studied.
Some researchers also use samples directly from ovarian tumors or adjacent tissues to identify dysregulated proteins in EOC. When the Westlake team reviewed clinical data, they found that none of the current leading therapeutic targets were discovered from these methods, although some are in clinical trials.
So, while proteomics has not yet led to therapeutics, researchers hope that as the field and technology progress, so will options for patients with EOC.
The challenges and promises of biomarkers
Biomarkers, typically proteins detected in patients’ blood or tissue, hold significant potential to help clinicians practice medicine more effectively. They can be used to diagnose and monitor disease progression, identify targets for drug development, monitor treatment effectiveness and predict which patients are at risk for drug side effects. Despite the promise of their utility, however, researchers have seen limited success and faced many challenges in bringing biomarkers to the clinic.
In a recent article in the journal Molecular & Cellular Proteomics, Jakob Bader and colleagues at the Max Planck Institute described challenges, technological advancements and success stories related to using mass spectrometry to identify biomarkers from body fluid samples. Many of the challenges were due to technological difficulties in high-throughput sample processing.
Researchers now address many of the challenges of biomarker discovery by using automation and simplified and improved protocols. Due to the many technological improvements the authors outline, it is possible to use larger cohorts of patients for better biomarker identification. This could mean the future is here, and with it, hopefully, the discovery of better biomarkers.
New technique helps detect protease cleavage
Proteases are enzymes responsible for cleaving proteins into smaller polypeptide chains or individual amino acids. This cleaving can modulate specific proteins and regulate key cellular processes. One such process is apoptosis, a form of programmed cell death, where various proteases are instrumental in initiating the breakdown of cellular protein components.
Proteases are important enzymes, and researchers need to be able to catalog their targets when studying the proteome. However, using mass spectrometry, or MS, to identify protein fragments generated by proteases is difficult.
To address this, Rawad Hanna and colleagues at the Technion–Israel Institute of Technology developed a new technique called LysN amino terminal enrichment, or LATE. This technique combines digestion with LysN and isotope labeling of the N-terminal end of a protein to better identify cleaved proteins in MS analysis.
The researchers recently described this work in the journal Molecular & Cellular Proteomics. By using LATE, they were able to identify novel targets of cleavage by caspase-3, a protease involved in apoptosis. They also discovered cross-talk between caspase-3 cleavage and N-terminal acetylation.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

