The promise of unknown unknowns
“There are also unknown unknowns — the ones we don’t know we don’t know. And if one looks throughout the history of our country and other free countries, it is the latter category that tend to be the difficult ones.” — Donald Rumsfeld
For soldiers and politicians, it is indeed the unknown unknowns that are difficult. However, as researchers, we can rejoice that, despite the difficulty, these are what make our chosen pursuit worthwhile.
Two essential fatty acids derived from plants are linoleic (18:2) and alpha-linolenic (18:3) acids. These dietary fatty acids are precursors of the omega-6 and omega-3 fatty acids popularized in the press. In addition to their importance for human health, these lipids also compromise oxidative stability of vegetable oils, and removing them results in the production of undesirable (and now banned) trans fats. Given the importance of these lipids, their synthesis in plants has gained considerable attention.
In oil-accumulating cells of plant seeds, linoleic and alpha-linolenic acids (referred to here on out as 18:2 and 18:3, respectively) are generated by the desaturation of oleic acid (18:1) catalyzed by fatty acid desaturase, or FAD, enzymes FAD2 and FAD3. For FAD2 and FAD3 to work, oleic acid must be incorporated into the membrane lipid phosphatidylcholine. How 18:1 is incorporated into phosphatidylcholine has been debated.
In earlier models, 18:1 was known to be incorporated into phosphatidylcholine by one of two routes: direct incorporation from 18:1-CoA catalyzed by acyl-CoA:lyso-phosphatidylcholine acyltransferase, known as LPCAT for short, or incorporation into diacylglycerol followed by conversion into phosphatidylcholine by CDP-choline:diacylglycerol cholinephosphotransferase, or CPT. It also has been proposed that reversibility of the CPT reaction would provide one mechanism for the production of polyunsaturated diacylglycerol for the synthesis of triacylglycerols containing 18:2 and 18:3.
During our earlier studies, we isolated a mutant of the model plant Arabidopsis with increased total levels of 18:1 and reduced total levels 18:2 and 18:3 in its seed oil. This mutant, rod1 (short for reduced oleate desaturation 1), had substantially reduced levels of 18:2 and 18:3 in both triacylglycerol and the immediate precursor diacylglycerol relative to wild-type Arabidopsis.
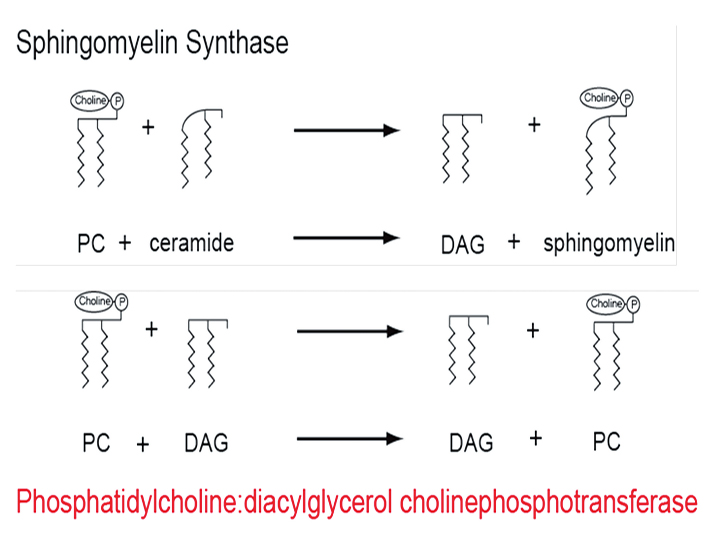 The known reaction between phosphatidylcholine and ceramide (top) provided the clue to identify a novel reaction in lipid synthesis (bottom) catalyzed by phosphatidylcholine:diacylglycerol cholinephosphotransferase.
The known reaction between phosphatidylcholine and ceramide (top) provided the clue to identify a novel reaction in lipid synthesis (bottom) catalyzed by phosphatidylcholine:diacylglycerol cholinephosphotransferase.
Surprisingly, however, phosphatidylcholine in the mutants contained increased 18:2 and 18:3. These data suggested to us that the rod1 mutation reduces transfer of 18:1 into phosphatidylcholine for desaturation but not the desaturation reaction itself. The known unknowns, lyso-PC acyltransferase and CPT, were eliminated from consideration by 14C-glycerol radiolabeling experiments and by sequencing the two CPT genes from rod1 and wild-type plants.
So what is ROD1? When the Arabidopsis genome sequence was completed in 2000, the puzzle of ROD1’s identity seemed to become a manageable task of identifying the gene locus by map-based cloning. However, when we completed this task and identified ROD1 as At3g15820, the encoded protein was annotated as a phosphatidic acid phosphatase-related protein. This made no sense in the context of our knowledge of the pathways of triacylglycerol synthesis or the fatty acid composition of rod1 seeds. Furthermore, assays of recombinant ROD1 protein found no detectable phosphatidic acid phosphatase activity.
Time for a Hail Mary pass? The geneticist’s equivalent of this desperate football strategy is a position-specific iterative basic local alignment search tool, or PSI-BLAST, search of protein databases. As in football, this approach most likely will get you in a mess, but there is always the chance that it will produce a winning touchdown.
Far down in the fourth iteration of our PSI-BLAST search (at entry No. 67) was a weak hit to a mammalian phosphatidylcholine:ceramide cholinephosphotransferase. This enzyme catalyzes transfer of the phosphocholine headgroup from phosphatidylcholine to ceramide, generating sphingomyelin and diacylglycerol.
Plants do not contain sphingomyelin, but the structure of ceramide is analogous in some respects to that of diacylglycerol, suggesting to us that ROD1 might catalyze the transfer of phosphocholine from 18:2/18:3-containing phosphatidylcholine to 18:1-containing diacylglycerol (see figure).
Indeed, assays of recombinant ROD1 confirmed this activity, revealing our discovery of a new enzyme of lipid metabolism, phosphatidylcholine:diacylglycerol cholinephosphotransferase. Called PDCT for short, this enzyme is required for efficient synthesis of 18:2 and 18:3, as well as some other fatty acids, during triacylglycerol accumulation in seeds. ROD1 also is expressed in vegetative tissues of plants.
Although the rod1 mutation does not result in substantial changes in root or leaf fatty-acid composition, it is possible that PDCT has a role in lipid homeostasis or remodeling of membrane lipids in response to temperature changes or other environmental perturbations.
ROD1 homologues are present in many plants, but no readily identifiable homologues are present in animals; however, the family of human proteins related to sphingomyelin synthase includes eight proteins of unknown function. Thus, it remains possible that PDCT will be found to play a role in lipid metabolism in animals as well.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

