JBC: A promiscuous inhibitor uncovers cancer drug targets
When signaling pathways within cells are triggered, proteins activate like a row of tumbling dominoes until the final protein influences some cellular function. In some tumors, multiple signaling pathways drive cell growth and survival; if one pathway ceases activity, another could continue driving cancerous behavior.
Several clinically approved kinase inhibitors originally were designed to block the function of individual signaling kinase proteins. Scientists now know that some of these inhibitors indiscriminately disrupt numerous proteins, which may allow them to kill certain tumors but also may inhibit some proteins unnecessarily, eliciting adverse effects.
To pinpoint the most therapeutically relevant kinase targets in cancer cells, a group of researchers at Harvard Medical School and the Dana Farber Cancer Institute in Boston, led by Nathanael Gray, has developed a method that exploits the multitargeted nature of a chemical inhibitor.
In a study published in the Journal of Biological Chemistry, Gray and colleagues identified key molecules that support the survival of a specific type of lung cancer. By analyzing how these cells respond to a cancer-killing kinase inhibitor with numerous targets, they showed that the anti-cancer effects likely were elicited by simultaneous inhibition of two signaling pathways. This approach could lead to development of cancer drugs that attack only the right targets.
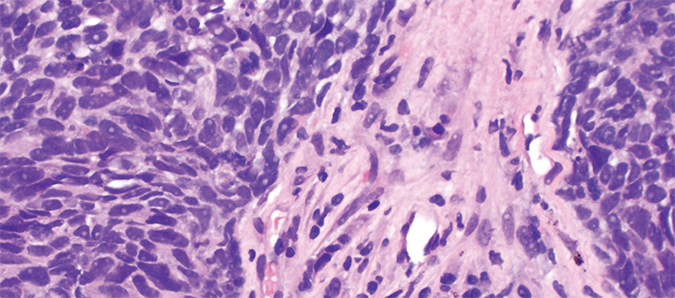 A micrograph of a non–small cell lung carcinoma shows nuclei in purple and cytoplasm in pink.Librepath/Wikimedia CommonsWhile studying various kinase inhibitors, Gray and his lab identified one, known as SM1-71, which binds to dozens of kinases, some of which support cell survival and growth.
A micrograph of a non–small cell lung carcinoma shows nuclei in purple and cytoplasm in pink.Librepath/Wikimedia CommonsWhile studying various kinase inhibitors, Gray and his lab identified one, known as SM1-71, which binds to dozens of kinases, some of which support cell survival and growth.
“It was sort of like a stick of dynamite and really could hit a lot of different targets,” Gray said.
In the JBC study, the researchers exposed several cancer cell types to SM1-71 and found that the drug was highly toxic to a lung cancer cell line with a mutation that activates many signaling pathways that drive cell growth. The inhibitor’s ability to kill these mutated cells suggested that targets in several pathways were being hit, Gray said.
To discover which of the many buttons pushed by SM1-71 elicited its anti-cancer effects, the researchers used Western blotting to narrow down which signaling proteins related to survival and growth were being blocked in cancer cells, revealing that proteins in two critical pathways were inhibited.
The authors then applied various kinase inhibitors to see if inhibiting any combination of the proteins in these pathways would replicate the cancer-killing effects of SM1-71. In the end, inhibiting MEK1/2 and IGF1R/INSR proteins at the same time demonstrated similar effects, suggesting that these are crucial targets in this lung cancer line, Gray said.
SM1-71 is likely not viable in humans because it binds to too many proteins and could cause collateral damage, Gray said. But uncovering its most important targets within specific pathways is valuable for designing drugs that can shut down multiple signaling pathways, which is necessary in some tumors.
“The next step would be to try to preserve the efficacy-driving targets while getting rid of targets that may be contributing to the toxicology,” Gray said.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

