From the journals: February 2018
We offer a selection of recent papers on a variety of topics from the Journal of Biological Chemistry, the Journal of Lipid Research and Molecular & Cellular Proteomics.
Two amino acids determine cells’ response to warming
 Drosophila, photographed in Dar es Salaam, Tanzania.Courtesy of Muhammad Mahdi Karim/Wikimedia Commons
Drosophila, photographed in Dar es Salaam, Tanzania.Courtesy of Muhammad Mahdi Karim/Wikimedia Commons
As the climate changes, we will have to adjust at many levels. In fruit flies, an enzyme called DESAT1, a delta-9-desaturase, is a key player in responses to temperature changes at both the cellular level, where it influences membrane fluidity by synthesizing monounsaturated fatty acids, and at the whole-organism level, where it controls temperature-influenced behaviors like mating.
Delta-9-desaturases are found in all organisms, from bacteria to humans. DESAT1 in fruit flies is transcribed at a constant rate, but the rate at which the protein degrades determines its overall expression level. Masato Umeda’s research team at Kyoto University was interested in understanding how the degradation of DESAT1 is regulated in the fruit fly Drosophila melanogaster in order to better understand thermal regulation in diverse organisms. The results of their research were reported in the Journal of Biological Chemistry.
The team discovered that DESAT1 is degraded by calpains, calcium-dependent cysteine proteases, in the presence of unsaturated fatty acids and that this degradation is enabled by a diproline motif at DESAT1’s N-terminus.
“It was surprising that it is regulated by only two amino acids,” Umeda said. “A very small portion is responsible for expression of the protein.”
Now, the team is going back a step further in the pathway leading to DESAT1 degradation, trying to understand how temperature changes are sensed inside the cell to lead to changes in fatty acid composition and consequent effects on the whole organism.
“Our ongoing project aims to elucidate how cells recognize temperature or cellular events caused by the temperature changes and control the expression of DESAT1 to adapt the cellular processes and behaviors,” Umeda said. “We hope our study will lead to an alternative approach that could potentially circumvent the biological crisis due to global warming.”
— Sasha Mushegian
Metabolites predict insulin resistance
Insulin resistance leading to high blood sugar, a hallmark of pre-diabetes, is a complex condition with many genetic and environmental determinants. Jacqueline Stöckli and colleagues from the University of Sydney carried out a comprehensive machine learning-enabled metabolomic analysis of three mouse strains fed high-fat or standard diets. The analysis showed that, despite individual environmental and genetic variation, a combination of three unrelated metabolites (C22:1-CoA, C2-carnitine and C16-ceramide) together formed an accurate signature for predicting insulin resistance. The study, published in the Journal of Biological Chemistry, is a step forward in creating precision medicine approaches to diabetes prevention and treatment.
Unclogging the plaques
Heart disease kills 18 million people in the world every year. The underlying pathophysiology of heart disease is often atherosclerosis, a progressive buildup of plaques within the arteries. While some components of the atherosclerotic plaques have been studied, a comprehensive picture of the changes that occur during development of the plaques is still lacking. Investigators at the Max Planck Institute of Biochemistry led by Matthias Mann aimed to capture this picture by using a sensitive and in-depth mass spectrometry approach to profile the aortas of atherosclerotic mice over the development of the disease. The researchers quantified 5,117 proteins from this profiling strategy and found that 182 of them changed significantly in response to vessel maturation and plaque development. These include novel factors that were not known to be involved in atherosclerosis, including matrilin-2, peroxidasin and MAM domain-containing 2. The study, published in Molecular & Cellular Proteomics, provides the vascular disease research community with a valuable resource of time- and compartment-resolved proteomic changes during atherogenesis.
Fatty acid metabolism and the migration of endothelial cells
Endothelial cells lining blood vessels help keep what belongs in and what doesn’t belong out. These cells move around as needed — during, for example, development and wound healing. Abnormal endothelial cell migration is seen in various diseases, including rheumatoid arthritis, cancer and macular degeneration. Researchers suspect that manipulation of vascular endothelial cell migration could be a useful therapeutic approach for those and other conditions. The enzyme known as ACC, meanwhile, catalyzes the first and rate-limiting step in the biosynthesis of fatty acids. This enzyme, formally acetyl-CoA carboxylase, is of interest to researchers developing therapies for metabolic diseases such as obesity and diabetes and for certain cancers in which ACC is upregulated.
So what do vascular endothelial cells and ACC have to do with one another? That’s the question that motivated a new study published in the Journal of Lipid Research. A research team led by Robert Fürst of Goethe University found that, in fact, ACC regulates endothelial cell migration. The study also underscored that manipulating ACC may be a worthwhile avenue for treating cell migration-related diseases. The researchers used human umbilical vein endothelial cells and human microvascular endothelial cells for the study. They interfered with ACC activities by administering a compound known as soraphen A. Originally isolated from slime bacteria, soraphen A has been used by scientists as an ACC inhibitor for some time. In this case, soraphen A’s inhibition of ACC changed the lipid composition of the endothelial cell membranes and had other effects that ultimately prevented the cells from migrating as usual.
“Our findings show for the first time that (the enzyme) regulates endothelial filopodia formation and, most importantly, endothelial cell migration by rearranging the cells’ phospholipid composition,” the authors report. “The present study fills a gap in the literature by showing the relationship between the fatty acid metabolism and the migration of endothelial cells.” In addition, the team noted that soraphen A could be a useful chemical tool for studying fatty acid metabolism in endothelial cells.
Trypsin trips up
Alternative splicing has been shown to occur in up to 94 percent of human genes, resulting in expression of different splice forms for each gene. While this phenomenon is known to increase complexity at the transcriptome level, the effect of alternative splicing on the complexity at the proteomic level remains controversial. In a recent study in Molecular & Cellular Proteomics, researchers at Baylor College of Medicine and Vanderbilt University led by Bing Zhang showed why there has been a lack of evidence of exon–exon junction-spanning peptides in published proteomic data sets. They found that the ends of exons in the genome are enriched with codons coding for lysine and arginine amino acids. This occurrence could have hindered the detection of junction-spanning peptides, since most mass spectrometry experiments involve digesting proteins with the enzyme trypsin, which cuts peptide chains exclusively after arginine and lysine residues. The researchers further showed that combining chymotrypsin and trypsin during sample preparation increased the detection of junction-spanning peptides by 37 percent. The study demonstrates the importance of using complementary digestion schemes in research where identifying different splice forms of proteins is a priority.
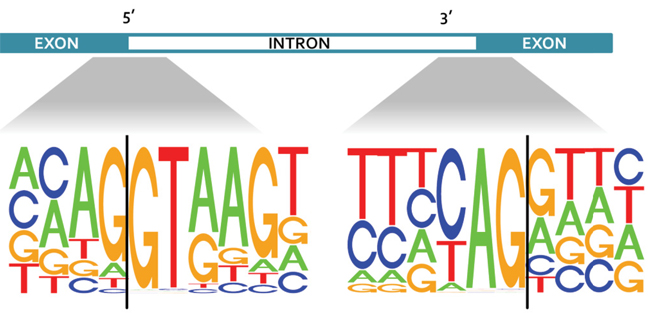 Consensus sequences on either side of the exon-intron boundaries for all coding genes in the human genome.Image courtesy of Bing Zhang
Consensus sequences on either side of the exon-intron boundaries for all coding genes in the human genome.Image courtesy of Bing ZhangMitochondrial signals through micro-RNAs
Communication between the nucleus and mitochondria is required for cells to function. While signaling from the nucleus to the mitochondria is most obvious, signals from the mitochondria to the nucleus are also crucial. In particular, mitochondrial-nuclear signaling regulates metabolic stress responses, and misregulation of this signaling has been observed in breast cancers. Trevor Carden and colleagues at the University of Alabama showed the first evidence of a micro-RNA’s role in mitochondrial-nuclear signaling and tumor suppression. The expression of micro-RNA miR-663 was induced by mitochondrial reactive oxygen species production, an apoptotic signal and oxidative stress response, and miR-663 induced the expression of nuclear-encoded mitochondrial respiratory chain subunits. Correlative and experimental observations of breast cancer tumors also suggested that miR-663 has a tumor-suppressive effect. These observations, published in the Journal of Biological Chemistry, suggest that micro-RNAs could be important players in mitochondrial-nuclear signaling in normal and cancerous cells.
 This illustration shows two views of a nicotinic acetylcholine receptor.Courtesy of Ataly/Wikimedia Commons
This illustration shows two views of a nicotinic acetylcholine receptor.Courtesy of Ataly/Wikimedia Commons
How nicotine affects receptor localization
Nicotinic acetylcholine receptors, or nAChRs, are the main receptors in neuromuscular junctions. Observations suggest that nicotine exposure affects the ratio of these receptors on the surface versus those in the endoplasmic reticulum of the cell and thus the cells’ sensitivity to these receptors’ natural ligands. To understand how nicotine affects nAChR localization, Ashley M. Fox–Loe and colleagues at the University of Kentucky developed sensitive, organelle-specific single-molecule imaging methods to quantify precisely changes in the stoichiometry of nicotinic receptors. Publishing the research in the Journal of Biological Chemistry, the team concluded that nicotine exposure affects receptor distribution through combined effects on both receptor assembly and trafficking, which eventually could help explain the effects of drugs like nicotine on neuromuscular functioning.
Investigating the genetic basis of bile acid diarrhea
Bile acid malabsorption, which can accompany gastrointestinal disease or manifest alone, causes chronic diarrhea. Primary bile acid diarrhea, or BAD, in humans has been shown to be related to the hormone known as fibroblast growth factor 19. FGF19 is secreted in the intestine and downregulates the synthesis of bile acids in the liver. Researchers at the David Geffen School of Medicine at the University of California, Los Angeles, recently reported in the Journal of Lipid Research that they have identified a gene variant that affects FGF19 secretion and thus may affect bile acid metabolism in both sickness and health. Led by Karen Reue, the research team first used a mouse model that they had identified previously as having a spontaneous deletion in the Diet1 gene. A deficiency in Diet1 results in reduced FGF15 (the mouse equivalent of FGF19). The team characterized the effects of the gene deletion and found that the mice had symptoms similar to those experienced by patients with BAD. Next, they looked at how Diet1 expression in more than six dozen mouse strains correlates with the expression of FGFF15 and bile acid levels. Finally, they zeroed in on a variant in the DIET1 gene in humans that influences FGF19 secretion from cultured cells and is associated with FGF19 levels in BAD cases. “Our findings raise the possibility that the (variant) could contribute to the seven-fold variation in plasma FGF19 levels that have been observed in healthy individuals,” the authors wrote. “Resequencing DIET1 in additional subjects will be required to reveal whether DIET1 rare mutations or additional common variants contribute to variations in bile acid metabolism.
Do-it-yourself custom chromatin
In the nucleus, DNA is packaged with proteins in a form called chromatin. Chromatin’s complex structure regulates gene expression, DNA repair and epigenetic inheritance. Building customized chromatin for molecular studies remains a challenge because of its multidimensionality. Mai T. Khuong and colleagues at the University of California, San Diego, developed a method to synthesize chromatin in vitro using histone chaperone proteins and an adenosine triphosphate-driven motor protein derived by a simple purification from Drosophila, template DNA and histones from the researchers’ species of choice, and ATP. This simplified system then was used to assemble chromatin with desired DNA sequences and proteins from diverse organisms. The researchers hope that this method, published in the Journal of Biological Chemistry, becomes a user-friendly approach for a variety of chromatin-related experiments.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

.jpg?lang=en-US&width=300&height=300&ext=.jpg)

