A very delicate balance
Earlier this year, the U.S. Food and Drug Administration approved the drug Aduhelm for Alzheimer’s disease. Its manufacturers say Aduhelm blocks aggregation of the protein amyloid beta in the brain. But evidence that the drug can help patients requires a very hopeful eye; clinical studies were so inconclusive that the agency’s advisory panel recommended against clearing the drug for marketing to patients. FDA leaders’ willingness to override that recommendation speaks to patients’ unmet need.
Whether the symptoms are the loss of mental clarity due to Alzheimer’s disease, the loss of physical control caused by Lou Gehrig’s or Parkinson’s disease, or another pattern of infirmity, neurodegenerative disease is a terrible diagnosis. Treatment options are limited, often restricted to managing symptoms with no effect on the slow advance of the underlying pathology. These diseases are estimated to affect about 6.5 million Americans today, and, as the population ages, that number is expected to rise.
Many neurodegenerative diseases share a feature: accumulation of misfolded proteins that neurons struggle to get rid of. Researchers are testing many possible ways to slow or reverse this aggregation, including using monoclonal antibodies such as Aduhelm that bind the offending proteins, reducing the activity of enzymes that produce them or boosting pathways that clear them. A few labs are investigating the possibility that a little-known enzyme called PIKfyve might offer another way to alter the course of aggregation and therefore disease. PIKfyve has an obscure role in cell biology, generating phospholipid species that are a vanishing minority of all the lipids in the cell. Yet the kinase seems to act as a gatekeeper to the lysosome, a key organelle in cellular waste disposal.
PIKfyve has been receiving attention from disparate sources recently, as the gene that encodes it turns up repeatedly in screens for genes affecting neurodegenerative disease. Researchers have found roles for the kinase in the development of Alzheimer’s, Parkinson’s and amyotrophic lateral sclerosis, or ALS — and they’re looking for ways to target it. The enzyme maintains a delicate homeostatic balance, and researchers in academic and corporate laboratories are eager to determine whether it can be manipulated safely.
But some researchers argue that we don’t know enough about PIKfyve to drug it successfully and that cutting off access to the lysosome could do more harm than good. The complex discussion is a case study in why it’s so difficult to develop new treatments for neurodegenerative disease.
Seeds and lysosomes
Belgian graduate student Alberto Soares wasn’t looking for lysosome regulators in 2015 when he started a study of cell-to-cell transmission of tau fibrils in Alzheimer’s disease, but the lysosome turns out to be important for how fibrils spread.
Fibrils, tough and durable structures formed when misfolded monomers of the protein tau glom together, make up the intracellular tangles that appear in brains with Alzheimer’s. The tangles spread from one part of the brain to another as the disease progresses, following known routes of neuronal connections. Researchers believe that misfolding of tau may be transmissible — that a misfolded copy of tau can, upon encountering a properly folded peer, propel it into the misfolded conformation. The disease-causing proteins called prions are well known to behave this way.
In Alzheimer’s disease, tau and amyloid beta accumulate; other diseases have been linked to aggregates of other proteins that also seem to spread. The model, sometimes called proteopathic seeding, is an increasingly prevalent and well-supported hypothesis.
Researchers still are investigating exactly how clumps of aggregated protein kill neurons — whether by blocking normal function or through some directly toxic pathway. The proteopathic seeding model also raises questions about how a fibril that starts inside one cell can gain access to properly folded proteins inside another.
Wim Annaert, a group leader at the Katholieke Universiteit Leuven in Belgium, is Soares’ research co-mentor. “If you can understand the mechanism of spreading and you can interfere with it,” Annaert said, “you might start looking at disease-modifying drugs that delay or even stop spreading.”
Annaert’s lab and collaborators in a lab at Janssen, the research arm of Johnson & Johnson, set out to understand whether a protein called clathrin, which is involved in endocytosis, helped to pick up the problem fibrils and introduce them into the cytoplasm. They found that it did not — but by the time they published early this year in the Journal of Biological Chemistry, that was almost an afterthought, eclipsed by something they came across accidentally.
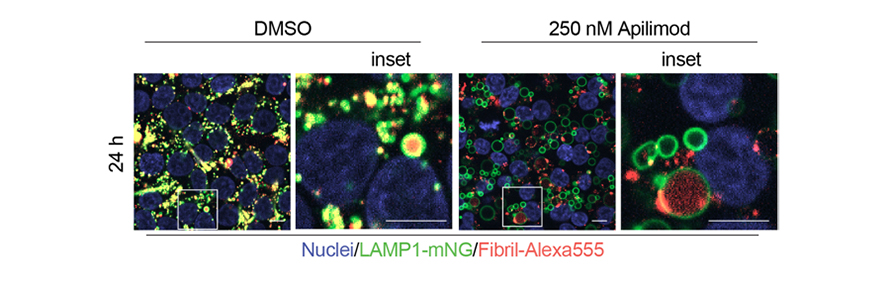
After adding an inhibitor to a protein downstream of endocytosis, Soares noticed different results depending on the dye he had used to label fibrils. Pursuing the discrepancy, he found that inhibition of the kinase PIKfyve prevents seeds a neuron has picked up from reaching the lysosome. This prevented fibrils tagged with acid-sensitive dyes from fluorescing. More importantly, the team found, it stopped fibril seeds from spreading.
To touch off a misfolding process throughout the cell, misfolded tau proteins first must gain access to the cytoplasm. To do that, they have to pass through the lysosome, which functions like the stomach of a cell: more acidic than the cytoplasm at large, it is filled with enzymes that break down complex molecules.
Proteopathic seeds give the lysosome a bad case of indigestion. When they’re delivered, for reasons that researchers do not yet understand, lysosomes tend to burst, letting seeds infiltrate the cytoplasm.
Louis De Muynck is a Janssen scientist who co-mentored Soares. “If you prevent the seed itself from ending up in the lysosome, you get less aggregation,” De Muynck said. “It’s not every day that we stumble upon these nice observations.”
PIKfyve, a lipid kinase, helps to give lysosomes and late endosomes their unique identity within the cell. Without the phosphoinositide species PIKfyve generates, endosomes and autophagosomes fail to fuse with lysosomes. And if tau fibrils never reach the lysosome, they never can escape from it.
The phosphoinositide code
Organelles are many and motley, filled with contents that differ from one compartment to the next. But from the cytoplasmic side, those components are obscured by a membrane wrapping. You might imagine the membranes of different organelles as closed doors to mysterious rooms.
Almost like paint color, the composition of each membrane can convey information about what’s behind each door. Subcellular membranes are composed of lipids that determine their fluidity, acidity, curvature and surface properties. Through different binding domains, lipids also help to determine which proteins are recruited to an organelle’s surface.
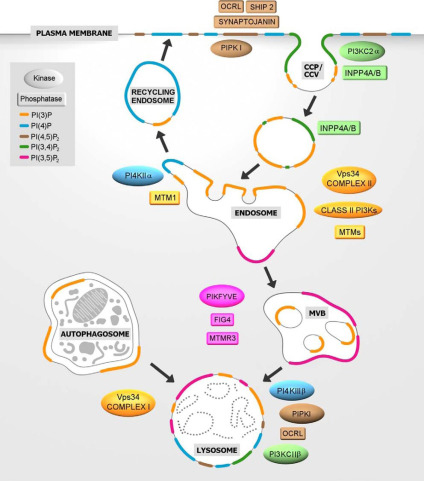
the distribution of phosphorylated phosphoinositides on membranes in the
endolysosomal system, and the enzymes that produce them.
Phosphatidylinositol, or PI, lipids are particularly important in giving each organelle membrane its character. Their head group, inositol, is unusually large, a sugar that can be phosphorylated at any — or all — of three hydroxyl groups. The combinations give a total of seven different phosphorylated species. Phosphorylated PIs are a small but influential subset of a cell’s total lipid species. Each is thought to be enriched at some stages of the endomembrane system, the network of organelles that starts with endocytosis and often terminates in the lysosome.
According to Scott Emr, a Cornell University professor who studies the roles phosphorylated PIs, or PIPs, play in membrane trafficking, it has become clear that the so-called organelle lipid code, in which one PIP species is enough to give a membrane its identity, is not an absolute. “That is, you don’t only find one PIP species on each organelle. However, each PIP is highly enriched on specific organelles,” he said, adding, “This preference in quantity on different membranes still supports the lipid code model.”
PIKfyve plays a key role in encrypting this code as the only kinase that can make the phosphatidylinositol species PI(3,5)P2, which is phosphorylated on the third and fifth positions of inositol. It also makes — whether directly or indirectly is hotly debated — the lipid PI5P. (See “PIKfyve pleiotropy.”)
Lois Weisman, a University of Michigan professor who has studied PIKfyve and its yeast homolog, Fab1, for decades, compared PIKfyve to the metabolic enzyme mTor, a well-studied protein kinase that conducts a metabolic symphony from its perch on the surface of the lysosome. PIKfyve, she said, regulates just as many processes. “Why PIKfyve isn’t as famous has nothing to do with its importance, in my opinion. It’s because measuring PI(3,5)P2 is next to impossible.”
PIKfyve pleiotropy
Some questions about PIKfyve – such as which lipids it produces and how those lipids are recognized – are still contentious topics.
READ MORESuch measurements, which rely on high-pressure liquid chromatography, are finicky and don’t give much information about cellular location. Thus far, efforts to develop a fluorescent sensor to visualize PI(3,5)P2 in cells have produced probes that probably bind other lipid species too.
Still, you don’t need a biosensor to see the most dramatic result of PIKfyve inhibition: enormous, numerous lysosomes.
Assia Shisheva, who identified and named PIKfyve in human cells soon after Emr and colleagues described its yeast homolog, also has worked on the kinase for many years. Shisheva, a professor at Wayne State University, was among the first to generate and express a mutant of the kinase that lacked enzymatic activity. “Initially when we were looking at these cells, we were saying, ‘Wait a minute. Why are the cells having this (appearance) — kind of like they’re taking a bubble bath? Is it something that we screwed up, the serum or something?”
But the cells expressing the wild-type enzyme were perfectly healthy, and the result appeared reliably in subsequent trials. The phenotype Shisheva observed was real; when PIKfyve was blocked, the cells gradually filled with vesicles that looked a little like soap bubbles.
Some effect of PIKfyve inhibition — whether the absence of PI(3,5)P2 or PI5P, or perhaps an excess of PI3P — closed the door to the lysosome. And strangely, when Shisheva and other researchers knocked out a phosphatase called Fig4 that reverses PIKfyve activity, they saw the same effect.
A marriage of opposites
Karin Reinisch, a Yale structural biologist who recently published on the structure of PIKfyve and its binding partners, said she was drawn to study the enzyme in part because paradoxes abound in the literature about it.
For starters, biochemists gradually had established that PIKfyve and the phosphatase Fig4 travel in tandem in the cell along with a third protein, Vac14. Given that the two enzymes have opposite effects, the observation didn’t make much sense. Reinisch said, “Nobody understood how you could have these two things in the same complex — how come they’re not just burning up ATP?”
In addition, it was unclear why knocking out Fig4 activity had the same effect as knocking out PIKfyve. “Why would you need to have phosphatase activity to have an active kinase?” Reinisch said. “Nobody understood that either.”

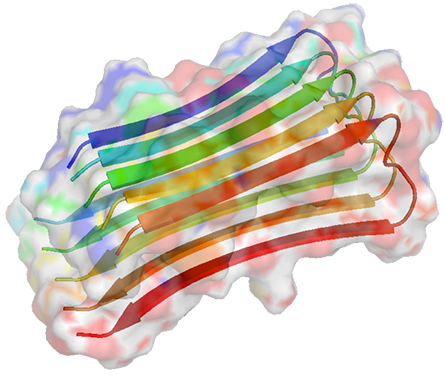
PI(3)P (bottom right) to produce PI(3,5)P2 (bottom left). Fig4, a phosphatase, removes that phosphate group.
Last year, researchers in Reinisch’s lab reported on a structure of the complex that helped clarify matters a little. The tentpoles of the complex are made of a star-shaped pentamer of the protein Vac14 cupped like a hand over the membrane. This five-pointed framework binds one copy of PIKfyve and one of Fig4. Based on the shape of the complex, Reinisch said, it’s likely that only one enzyme’s active site can gain access to the membrane at any moment, preventing wanton waste of ATP.
While PIKfyve and Fig4 have opposing effects on the phosphorylation of their lipid substrate, each one also acts on proteins — and here, they help each other out. When PIKfyve is active, it autophosphorylates, inhibiting its own lipid kinase activity. Meanwhile, Fig4 can dephosphorylate the kinase, which increases its activity.
“The interactions between the three proteins are very, very complex,” Shisheva said, pointing out that her lab previously had described the counterintuitive effect of Fig4 on PIKfyve activity.
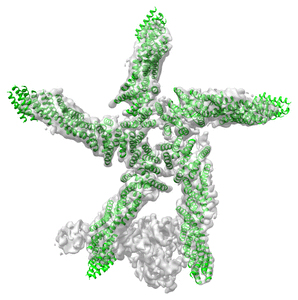
According to Weisman, the complex, almost Rube Goldbergian interaction could give the cell numerous ways to open or limit access to the lysosome. “I think the cell needs a way to acutely turn this pathway on and off,” she said, noting that in yeast cells under acute osmotic stress, PI(3,5)P2 concentration surges rapidly and then drops off as the cells adapt to their new conditions.
Reinisch said the structure leaves numerous questions unanswered. PIKfyve is phosphorylated at many additional sites, and the effects of those other modifications are unclear. In addition, it’s hard to tell from the cryo-electron microscopy structure exactly how PIKfye inhibitors might fit into the protein’s structure.
“We’ve been contacted by several people (to ask) could we crystallize the kinase with the inhibitor,” Reinisch said. Thus far, that has been impossible because “the molecule is very big, with many domains, and it’s breathing and it’s moving around, so we can’t get to a higher resolution.”
PIKfyve and neurodegeneration
The researchers asking Reinisch about the inhibitor probably are intrigued by the number of independent high-throughput screens, using different methodologies and investigating different diseases, that recently have pointed to PIKfyve as a promising drug target.
In addition to the JBC study on tau, researchers have identified PIKfyve in screens for regulators that control the aggregation of alpha-synuclein in Parkinson’s disease, modifiers of the deterioration of neurons with mutations in the ALS-related protein C9Orf72, and driver mutations in thousands of patients with neurodegenerative disorders. In addition, Fig4 mutations have been known for decades to link to a subtype of the degenerative nerve disorder Charcot–Marie-Tooth disease.
“Not that many people are interested in (PIKfyve), but, because these disparate groups are, it makes you think maybe something’s really there,” Weisman said of the emerging trend in the literature.
Stephanie See, a postdoc at the University of California, Berkeley, said, “I think it’s very possible that it’s sort of a universal mechanism … important for controlling the spread of many aggregating proteins.”
As a graduate student, See worked in Martin Kampmann’s lab at the University of California, San Francisco. In a recent preprint, she and colleagues reported that using a CRISPR loss-of-function screen to remove some enzymes in the phosphatidylinositol pathway could slow the spread of alpha-synuclein fibrils in human embryonic kidney cells. In follow-up experiments using various specific inhibitors for phosphatidylinositol-regulating enzymes, See found that a PIKfyve inhibitor had the most dramatic effect, reducing aggregation almost completely.
“From a therapeutic standpoint, that was really interesting for us,” See said. “It’s kind of easy to break a system. But if you can inhibit something and decrease the pathological phenotype, that’s really, really cool.”
Many researchers have considered attacking proteopathic diseases by turning up autophagy, the cell’s system for directing bulk waste to the lysosome. But inhibiting PIKfyve should do the opposite.
Justin Ichida’s lab at the University of Southern California posted a preprint in 2019 showing that in cultured neurons with mutations linked to ALS and frontotemporal dementia, inhibiting PIKfyve had a neuroprotective effect. “It was really puzzling when we first found it,” Ichida said. “Why would blocking something that blocks autophagy be helpful here? It was kind of counterintuitive.”
In the studies from Ichida’s and Kampmann’s labs, as in the JBC study that focused on tau, the bubble-bath phenotype that arises after inhibiting PIKfyve seems to be important in helping cells. “By blocking PIKfyve, you trap the fibrils in early endosome compartments,” See said. “They can’t get to the place where they can break out of the endosome and cause problems.”
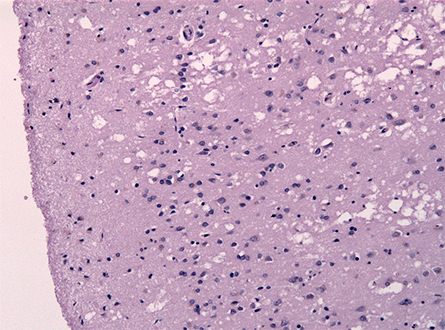
by prion disease.
But for some researchers, vacuole-filled cells have ominous implications. Asked about blocking PIKfyve to treat neurodegenerative disorders recently, Adriano Aguzzi, a neuropathologist at the University of Zurich, answered quickly and emphatically. “No, no, no. Don’t do that. It will kill you.”
Aguzzi calls the phenotype spongiosis. “It’s so characteristic and so unique that, as a pathologist, if you look at the brain and see spongiosis, you know within a fraction of a second that there is a prion disease.”
His lab reported in a paper published in July that after deliberate prion infection in cultured neurons or in mice, PIKfyve vanishes. “The moment the mice are really terminally sick, there is no PIKfyve detectable in the brain anymore,” Aguzzi said. “This was one of the very first times in my life where a result coming out of a blot genuinely surprised me.”
Following up the observation with mechanistic biochemistry, the researchers found a causal chain from an unfolded protein response to PIKfyve destabilization to the bubble-bath phenotype. In samples from a brain biobank, they observed that patients with more dramatic loss of PIKfyve had succumbed to prion disease earlier.
Weisman said the Aguzzi lab’s findings “broadly fit with our own earlier finding that mouse mutants deleted for Vac14 or Fig4, or with a hypomorphic mutation in PIKfyve, exhibit spongiform brains.”
Aguzzi is less sanguine than many colleagues about inhibiting PIKfyve to prevent the spread of proteopathy from one cell to the next. “I think it’s a very delicate equilibrium between PIKfyve and Fig4 and the amount of spongiosis,” he said, “and I would be super scared of touching that.”
Drugging PIKfyve
Despite the concerns, preliminary results from numerous studies of as-yet-untreatable diseases of protein aggregation have piqued drug developers’ interest in PIKfyve.
“There’s a lot going on in the PIKfyve therapeutic development world,” Ichida said. “Our company, Acurastem, is making an antisense therapeutic for PIKfyve; Verge Genomics is making a small-molecule inhibitor; AI Therapeutics is moving forward their small-molecule inhibitor; then there are other large companies that are trying to either develop their own programs or license other programs.”
The most advanced of these drug development programs, at AI Therapeutics, was conceived as a way to kill cancer cells, not to save neurons. (See “The many lives of apilimod.”) But the company announced early in 2020 that, while doing large-scale data mining, it had linked PIKfyve to ALS and that collaborators in Icihda’s and other academic labs had found that ALS-model neurons survive better in culture when treated with AI’s PIKFyve inhibitor, apilimod.
Apilimod already has made it through the first phase of drug development, clinical safety testing. While some study participants suffered nausea and vomiting at high doses, lower doses seemed to be reasonably well tolerated. Based on those safety results, Ichida and other scientists working in this area argue that there may be a therapeutic window — a dose that delivers benefits without harmful side effects — for PIKfyve inhibitors.
“Clearly, people can survive having been treated with this molecule, right?” Weisman said. “So either these vacuoles aren’t arising, or maybe they are being resolved somehow.”
The many lives of apilimod
The molecule apilimod wasn’t always recognized as a PIKfyve inhibitor. At first, researchers thought it might block cytokine signaling.
READ MOREPIKfyve optimists also cite genetic evidence. Although an embryo without PIKfyve will die very early in development, people can be born with just one functional copy of the gene. Those individuals seem to suffer no ill consequences except for a tendency to develop white flecks within their corneas that can mildly impair vision, and which upon examination turn out to be cells filled with swollen vacuoles.
Perhaps a drug could reduce PIKfyve activity enough to slow lysosome fusion, and therefore seed propagation, without halting it altogether. Ichida said that his company, Acurastem, has conducted but not yet published experiments applying its PIKfyve inhibitors to a mouse model of ALS. There, he said, “Vesicle enlargement doesn’t seem to be something that’s required for the efficacy or the therapeutic rescue.”
Or maybe vesicles could be cleared by a mechanism that doesn’t involve the lysosome. Ichida noted that even if vacuoles do develop, scientists have found that some cultured cells activate secretory autophagy when PIKFyve is blocked. He suggested that the phenomenon, which happens when autophagosomes blocked from the lysosome dump their contents into the extracellular space, could act as an escape valve for the extra vacuoles.
Like researchers at AI Therapeutics, scientists at Verge Genomics picked up on PIKfyve as a potential target based on mining genetic data from ALS patients. Irene Choi, a senior director and head of drug discovery at Verge, said her team has not observed abnormal vesicle formation in cells treated with their lead candidate, a small-molecule inhibitor of PIKfyve. “But of course, we weren’t specifically looking for that either, at that time. We are currently exploring what potential liabilities vacuole formation imposes,” Choi said. “What we do know is that, in our model systems, we don’t see an accelerated cell death; we actually see an improvement in cell death over time.”
Like Ichida, Choi said that the lack of neurological danger signals from clinical trials of apilimod, which has been tested in hundreds of patients, some for six months or longer, was an encouraging sign that PIKfyve inhibitors might be safe to use.
The whole drug-development industry runs on this kind of bet: being the first to stake out, and patent, a therapeutic area and then making sure over time that the treatment strategy is safe for patients and achieves the desired result. It’s part of the reason so many drug candidates fail before reaching patients.
What happens now?
Researchers are left with a puzzling combination of observations to sift through. Will low-grade inhibition of PIKfyve be a useful brake on uncontrolled spreading of pathological protein aggregates, or will it kill neurons by overfilling them with vacuoles? Is closing the door to the lysosome beneficial or harmful?
“I would like to believe that this is going to pan out,” Weisman said. Still, she acknowledged that she has her doubts. The mixed sentiment is widely held. No one is rooting for neurodegenerative diseases, but many potential mechanisms for therapy have failed over time.
Could blocking PIKfyve be therapeutic? “I think that remains to be seen,” Kampmann said. “The nice thing about the cell-based studies is we now know what to look at” during a transition into model organism studies. Kampmann added that his lab has found evidence that other phosphatidylinositol pathway enzymes are involved in the onset of neurodegenerative disease.
Even Aguzzi, the most emphatically skeptical researcher interviewed for this article, thinks studying PIKfyve could lead to useful knowledge about neurodegeneration. His lab has started to run CRISPR screens in search of modulators of PIKfyve activity that might influence vesicle trafficking.
Exploration in the pharmaceutical labs is likely to determine whether neurons in a living brain fill with vacuoles during sustained pharmaceutical inhibition of PIKfyve.
Even if PIKfyve inhibitors don’t prove to be a safe and effective way to block the spread of seeds, researchers still may be able to parlay knowledge about PIKfyve’s role into useful approaches to combat neurodegeneration. De Muynck, the Janssen scientist, said, “We’re keeping an eye open.”
PIKfyve pleiotropy
PIKfyve phosphorylates PI3P, generating PI(3,5)P2. On this much, everyone agrees.
In mammalian cells, the enzyme PIKfyve contributes to levels of PI5P, which is phosphorylated at just one location. But scientists have yet to reach consensus — and rumors fly about major meeting arguments — over whether PIKfyve generates PI5P directly, by phosphorylating PI, or indirectly, by raising the amount of PI(3,5)P2 that phosphatases can act on.
Whether direct or indirect, said University of Pittsburgh lipid biochemist Gerry Hammond, “The controversy comes from the fact that there are so many experimental observations that would agree with both models.”
In addition, PIKfyve activity may slightly lower the level of its abundant substrate PI3P, which could contribute its own effects to overall cell biology. As with many metabolic pathways, Hammond said that it’s very difficult to disentangle the lipids’ contributions when PIKfyve is inhibited.
Those effects, he added, depend on more than just the presence of specific phosphorylated lipid species. “There’s kind of two parts of the story,” Hammond said. “There’s the biochemistry to figure out the enzymes that are involved in making these lipids. But then there’s a class of effector proteins.”
“With PI(3,5)P2 we don’t have a very complete list yet of bona fide effector proteins,” Hammond added. It is likely that those effector proteins — whatever they are — govern many of the downstream effects of PIKfyve activity, which are thought to include lysosome acidification, fusion between endosomes and the lysosome, and perhaps also fission.
A similar mystery hovers around where, exactly, the lipids in question are most effective. “We do think that more than one compartment is affected by PIKfyve inhibition,” Martin Kampmann said. His lab and Lois Weisman’s have observed PIKfyve in cellular neighborhoods other than the lysosome.
“When people would inhibit something with apilimod, then they’d see these huge swollen lysosomes,” Weisman said. “That was the most striking phenotype. … But then, you have to wonder: is that all PIKfyve does?”
The many lives of apilimod
When the molecule now called apilimod first was reported in 2006, researchers believed it acted by inhibiting interleukin production in T cells; its inventors figured that this might be useful for the treatment of Crohn’s disease. It took several years before another pharmaceutical team reported that the molecule actually inhibits PIKfyve and that PI(3,5)P2 was an intermediate step in the T-cell signaling that leads to release of some interleukins.
In 2017, cancer researchers at AI Therapeutics reported that non-Hodgkin’s lymphoma cells are highly sensitive to PIKfyve inhibition, dying when exposed to a low concentration of apilimod. The cells, the authors concluded, were more dependent on autophagy and the downstream lysosome activity than ordinary cells. Partial data from a clinical trial testing apilimod for safety in people with non-Hodgkin’s lymphoma are available. Meanwhile, the company appears to have pivoted from using apilimod for oncology and now is testing it as a small-molecule antiviral against SARS-CoV-2 in phase 2 preliminary efficacy trials.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.



