With a zap of electricity, scientists write a message into bacterial DNA for the first time
Harris Wang and his team at Columbia University have harnessed the power of living bacteria to write binary code in the bacteria’s own DNA. This approach is the latest in a quest to use DNA to store digital data. Their study is the first, however, to make and store that DNA directly in living bacteria.
These efforts are a response to some experts’ concerns that the needs of a digital world are outpacing the supply of computer chips needed for traditional hard drives. By turning instead to bacteria, this new research reimagines DNA for a digital purpose.
DNA is an enticing storage molecule because it fits a wealth of information into a microscopic package. All of the complicated instructions that make you unique fit into a cell nucleus that’s approximately one-tenth the width of a single human hair. If you unwound and stretched out the DNA in a single cell, it would measure six feet long. That adds up to nearly 67 billion miles of DNA in your body alone. DNA can store petabytes of data per gram, making it one of the most information-dense materials on earth.
One significant hurdle has stood in the way of past attempts to use DNA as a storage molecule for digital data. Long messages require long strands of custom DNA, which are expensive to make with typical DNA synthesizer machines. Wang’s team overcame this hurdle by turning to a much cheaper DNA-making machine: live bacteria.
Wang’s research team developed what they called an “electrogenetic DNA writer” — an entirely new bacterial system that uses electrical signals to alter the DNA inside living bacteria. They did this by drawing inspiration from CRISPR genome editing. CRISPR utilizes a complex of specialized enzymes to insert short fragments of DNA into the bacteria’s genome. The fragments are inserted one right after the other. These fragments — known as “spacers” — have unique sequences that stand out from the rest of the bacteria’s DNA.
The research team recognized the potential to string these spacers together into a pattern that could encode a message. Instead of using a unique spacer for all 26 characters in the English alphabet, they used the two digits of binary code to simplify the system.
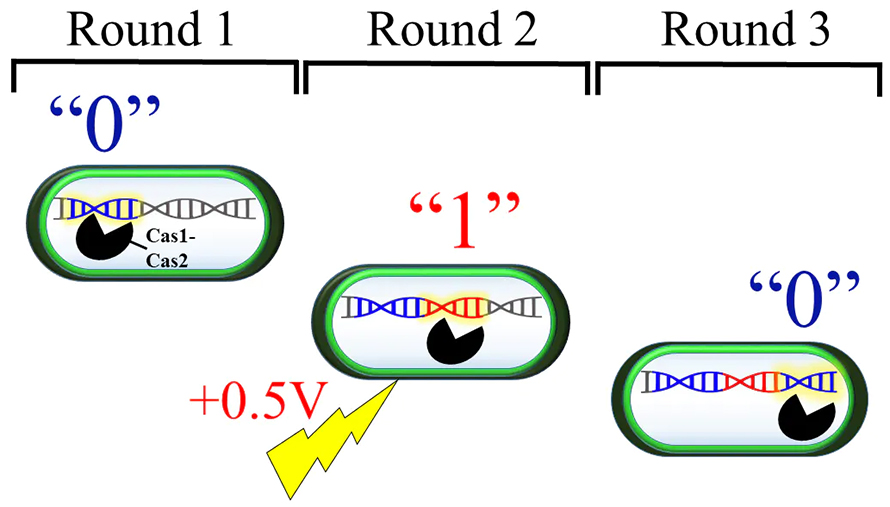
The team developed two types of spacers that stand for the “0s” and “1s” in binary code and controlled when each spacer type was inserted into the bacterial DNA with an electrical pulse. In the absence of an electrical signal the CRISPR complex inserted the spacer corresponding to a “0”, while a +0.5V electrical pulse set off a chain reaction that inserted the spacer corresponding a “1”. With this system the researchers were able to string together sequences of “0s” and “1s” to code the message “hello world!” directly into bacterial DNA.
This is not the first time that DNA has been used as a digital data storage device. Past attempts, however, required expensive DNA synthesizers to make custom strands of DNA. In 2012, one research group encoded an entire 53,426-word book in DNA. Less than a year later, another created custom DNA molecules that stored, among other things, all 154 of Shakespeare’s sonnets and Dr. Martin Luther King, Jr.’s “I Have a Dream” speech.
While these efforts proved that DNA could store unique messages, those long synthetic strands of DNA are too expensive for widespread implementation. We are on track globally to produce 163 zettabytes (or 163 trillion gigabytes) of data by 2025, meaning that any useful data storage solution must be inexpensive and scalable. With the help of one of the Earth’s earliest life forms, DNA data storage could be a cost-effective and realistic possibility.
This new bacterial system offers benefits beyond a reduction in cost. Living bacteria can shelter the DNA message in harsh environments that would kill unprotected DNA. When Wang’s team mixed DNA messages into commercial potting soil, DNA alone degraded after about six days while DNA stored in bacteria remained intact and readable. Scientists already have their eyes set on using radiation- and heat-resistant strains of bacteria in the future.
Don’t expect to store your family’s vacation photos in bacteria just yet, however. This technology still has a long way to go before it replaces your computer’s hard drive.
The error rate of these “living hard drives” is likely too high for commercial use with large amounts of data, even with the team’s best error correction model. The living bacteria also presented a problem. As bacteria divide, mutations to the DNA sequence can occur. In this study, the coded message lost accuracy after about 80 generations of bacteria, which proves particularly problematic for longer messages (in Escherichia coli, the bacteria used in the study, a generation in the wild is between 12-24 hours, but can be as little as 20 minutes in a nutrient-rich lab setting). So, while the successful encoding of “hello world!” proved that this technology can store short messages, a more robust data-storage solution will need to handle much longer messages.
Even with these drawbacks, “living hard drives” offer an exciting way forward. Many of the issues the researchers encountered could be addressed with additional engineering of the CRISPR system. Armed with the knowledge that coded messages hold up in natural environments — could future James Bond films feature secret messages hidden in muddy shoe prints?
This story originally appeared on Massive Science, an editorial partner site that publishes science stories by scientists. Subscribe to their newsletter to get even more science sent straight to you.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

