World Cancer Day
Today is World Cancer Day, an observance focused on understanding the causes and treatments of cancers, improving equitable access to healthcare, and advocating for policies to reduce the incidence of and improve treatment of cancer. Much progress has been made recently in understanding the development and progression, paving the way for new insights and therapies for cancer treatments.
What is cancer?
Cancer is a broad term used to address a collective set of diseases. Though cancers can occur in many tissue types and can have a diverse set of causes, all cancers share a particular set of characteristics.
First, cancers grow, or proliferate, uncontrollably. Normal cells take steps to regulate their proliferation. For example, as a cell divides, every cycle of cell growth and DNA replication causes limited DNA damage. When this DNA damage becomes too severe, a normal cell will exit the cell cycle and stop dividing, a state called senescence. Alternatively, if the DNA damage is irreparable and too severe, a normal cell will kill itself, a process called apoptosis. Cancer cells find ways to extend the lifetime of DNA and to suppress the signaling pathways that cause senescence or apoptosis. Consequently, cancer cells grow and divide almost indefinitely, ignoring or suppressing the usual signals to stop cell growth. A growth of cancer cells forms a tumor.
Cancer cells change as a result of their new growth programs. Because cancer cells are constantly dividing, they have an increased energy uptake and may cause nearby blood vessels to grow toward the cancer to increase the nutrients to the tumor. Since some cancer cells consume so much oxygen, cancer cells have to find ways to grow in the presence of less oxygen. Moreover, cancer cells reprogram their metabolism in order to accommodate their new energy demands. Since each round of cell division causes DNA damage and cancer cells, a cancer cell’s DNA can become unstable, and a cancer cell may exhibit aneuploidy, or an unusual number of chromosomes due to error-prone cell divisions. A cancer cell may also duplicate DNA regions containing genes favorable to growth and delete DNA regions corresponding to genes that would normally stop it from growing or spreading.
Finally, many cancers can spread across the body, or metastasize. To accomplish this, cancer cells undergo another round of reprogramming to break away from their tissue of origin and become more mobile. They then travel through the body (usually through the blood stream or lymphatic system), escaping the detection of the immune system, and reattach at a new site. At the new site, a new tumor can develop, and the tumor may cause an organ to fail and may lead to death.
In the United States, the most common cancers are breast cancers, lung cancers, prostate cancers, and colon cancers.
What are common causes of cancers?
Cancers have a number of causes, broadly defined into two categories: lifestyle causes and genetic causes.
A number of genetic changes are necessary for the initiation of cancer. In general, cancer cells have activated at least one mechanism to encourage inappropriate growth and have deactivated at least one mechanism that would have regulated and slowed its growth. One of the most commonly mutated genes in cancer is TP53, which codes for tumor protein 53 and is activated in response to DNA damage. The protein p53 causes cells to stop growing or to commit apoptosis, thus preventing cancers from developing. When mutations cause p53 to misform, it is generally inactivated, and loss of p53 function is thought to play a role in about 50% of many cancers. Another commonly mutated gene is RAS, which codes for a protein that activates central growth pathways within the cell. When RAS is mutated, it is constitutively activated; it cannot be turned off. Consequently, cells with RAS mutations are constantly stimulated to grow and can develop into cancer cells. Genes like p53 that usually slow the development of cancer are known as tumor suppressor genes, whereas genes like RAS that support cancer progression are known as oncogenes. Genes like these are commonly mutated, either to inactivate tumor suppressor genes or to hyperactivate oncogenes, in cancers.
Certain lifestyle choices can increase the chance of DNA mutation and consequently cause cancer. Smoking, excessive alcohol consumption, and other drugs can cause DNA damage in a cell that may turn it from a normal cell into a cancer cell. Viruses, such as human papilloma virus; excessive exposure to UV light, such as sunlight or tanning beds; and exposure to certain kinds of pollution or chemicals, such as cigarette smoke and lead, also can change a cell’s DNA and transform a normal cell into a cancer cell. An unhealthy diet and unhealthy lifestyle choices, such as obesity and lack of exercise, also have been linked to an increase likelihood of cancer. Finally, since cells accumulate DNA damage as they get older and divide, the likelihood of cancer increases with age.
What are active areas of cancer research?
Current cancer research is focused on identifying the proteins and genes responsible for cancer development and progression, understanding their role in cell division and cancer development, and finding methods to therapeutically treat cancer. All the characteristics of cancers are common research areas for understanding the progression and treatment of cancer.
Inhibiting tubulin
All cancer cells divide uncontrollably, so identifying drugs that can target and disable the cell-division machinery is a common mechanism of cancer treatment. Tubulin is one component of the mitotic spindle, the tubulin-based structure cells need to divide. Many common chemotherapies, including paclitaxel (brand name Taxol) and vinblastine, inhibit the function of tubulin, but these drugs come with severe side effects. In contrast, tirbanibulin, a tubulin inhibitor in clinical development, has shown significantly lower toxicity or side effects than other tubulin inhibitors. To understand why, Niu et al. in China studied the interaction between tubulin and tirbanibulin and reported in the Journal of Biological Chemistry in October that tirbanibulin binds reversibly to tubulin, something not shared by another tubulin inhibitor, colchicine. The authors suggest that this reversible binding may be the cause of the tirbanibulin’s low toxicity in human patients and may inform future clinical use of tirbanibulin.
AMPK in normal and cancerous cells
Cancer cells have unique metabolisms that can be targeted for chemotherapies. AMPK, an enzyme involved in cell metabolism, is an important protein for recognizing and responding to the energy demands within a cell. Bartel et al. in Germany reported in JBC in October that, when treated with the drug archazolid, AMPK levels increase in noncancer cells, but AMPK levels are high in cancer cells regardless of whether or not the cells are treated with archazolid. In noncancer cells, heightened AMPK levels protect the normal cells from archazolid, but in cancer cells, the protective effect of AMPK is lost and the cancer cells die. The authors suggest archazolid and drugs similar to it may be promising therapeutics, as they specifically target cancer cells and not normal cells.
USP3 and breast cancer
Identifying genes and proteins that are necessary for cancer cell progression is important for identifying targets for cancer treatment. KLF5 is a transcription factor overexpressed in breast cancer that promotes cell proliferation, survival, migration and tumorigenesis. Wu et al. discovered the protein USP3 stabilizes KLF5. Many proteins are degraded via ubiquitylation. In ubiquitin-mediated degradation, the proteins to be degraded are first tagged with many ubiquitin molecules. Next, the ubiquitylated proteins are recognized for degradation by the ubiquitin proteasome system. USP3 is a deubiquitylase, meaning it removes ubiquitin molecules from a protein to be degraded. Since the target protein is no longer ubiquitylated, it is no longer degraded. Decreasing the level of USP3 lead to slower cell growth and smaller tumor sizes in mouse models of breast cancer, suggesting USP3 may be a therapeutic target in the treatment of breast cancer. The researchers, based in China, reported their work in JBC in October.
Regulating NOTCH1
Similarly, understanding why cancer develops is important to design therapies for treatment. The protein NOTCH1 regulates a signaling pathway that can both suppress and encourage cancer growth, but what factors decide between the two fates is poorly understood. De Blasio et al. in Italy discovered mitotic kinase Plk1 as a potential link between these two opposing NOTCH1 functions. The authors found that Plk1 phosphorylates NOTCH1, and NOTCH1 is consequently degraded. The authors also demonstrated that, during periods of DNA damage, NOTCH1 levels increased and NOTCH1 arrested cells from dividing. Moreover, Plk1-mediated degradation of NOTCH1 was necessary for return to normal cycling after such a cell-cycle arrest. However, this induced-Plk1 activity also may contribute to inappropriate progression into mitosis and error-prone cell division. Thus, Plk1 may be one factor that regulates NOTCH1’s role in both stopping cell division due to DNA damage and in promoting error-prone cell division. They reported their work in JBC in October.
Engineering more effective CAR T-cells
Reprogramming a patient’s immune system to recognize cancer cells is an emerging form of cancer treatment termed CAR-T cell immunotherapy. For CAR T-cell therapy to be effective, the CAR T-cells need to stick or adhere to the site of the tumor so that the immune system can attack the cancer cells. Mondal et al. realized that they could engineer CAR T-cells to be more effective by modifying the CAR T-cells to express the carbohydrate sLEx, part of the protein complex used to adhere circulating blood cells to the walls of blood vessels. CAR T-cells expressing sLEx were “stickier” and, in mouse models, found with higher frequency in bone marrow than CAR T-cells not treated to express sLEx. These findings suggest that CAR T-cells can be targeted to certain tissue types by engineering the carbohydrates on the surface of the T-cells. The researchers, based in Massachusetts and Florida, reported their findings in JBC in October.
A better prostate cancer biomarker?
Early detection of cancer is an active field of cancer research, because early treatment leads to better clinical outcomes. Accurately detecting cancer usually depends on detecting the presence of one or more cancer-specific proteins or other biomarkers. Obtaining a tissue sample to test ideally should be noninvasive and easy to obtain, such as hair or saliva. Prostate-specific antigen (PSA) is a commonly used biomarker for prostate cancer. Many patients with prostate cancer have elevated levels of PSA. However, PSA is secreted by both normal and cancerous prostate cancer cells, and elevated PSA levels are associated with other diseases, not just prostate cancer, making PSA a moderately useful biomarker for prostate cancer. Drabovich et al. are among those researchers seeking better prostate cancer biomarkers. Using mass spectrometry, they looked at seminal fluids from patients with and without prostate cancer, identified the proteins present, and used a machine-learning algorithm to determine which proteins were good biomarkers for the presence of prostate cancer. This method revealed that the levels of the protein TGM4 is about three times higher in samples from those with prostate cancer than in those from patients without. Moreover, higher TGM4 levels could be used to identify prostate cancer in both normal and cancerous PSA-elevated patient samples. These results suggest that TGM4 levels can help doctors determine whether PSA levels are elevated because of the presence of prostate cancer or because of another condition. The results were published in the journal Molecular & Cellular Proteomics in September.
Search for oral cancer biomarkers continues
Oral cancers are usually identified visually, but they can look similar to other oral conditions, and some oral cancer patients are unable to open their mouths for a medical evaluation. Another challenge is the fact that researchers have so far be unable to identify biomarkers for certain oral cancers. Recently, however, Chu et al. analyzed saliva samples from healthy patients, patients with oral cancer, and patients with visually similar oral conditions and identified three biomarkers that could be used to distinguish the saliva of patients with oral cancer from healthy patients or patients with other oral conditions. The published their study in MCP in September.
Detecting ovarian cancer
Similarly, Rüttenhain et al. recently developed another mass spectrometry-based approach to identify biomarkers from the blood to improve the specificity of tests for detecting ovarian cancer. From this analysis, they identified a five-protein signature that, when combined with existing tests, improved the specificity of ovarian cancer detection. Their work was published in MCP in September.
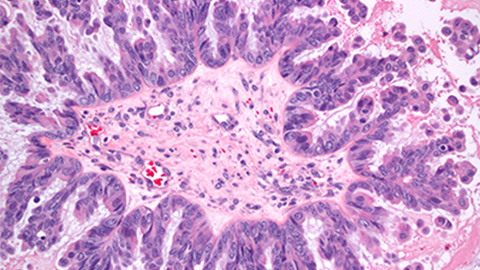
Catching ovarian cancer
when it’s curable
In the journal Molecular & Cellular Proteomics, researchers at Tel Aviv University reported a new test for ovarian cancer that outperforms previous tests. They hope it can be used to screen women who are genetically predisposed to the disease.
Overcoming resistance to treatments
Despite advances in the treatment of cancers, cancer cells can become resistant to the drugs used to treat them. In certain types of leukemias, cancer cells produce proteins that pump the chemotherapy out of the cell, lowering the effectiveness of the drug. To understand what other factors affect drug resistance, Kao et al. characterized leukemia cells resistant to DNA-damaging agents and found that they had increased concentrations of the sphingolipid ceramide. Using inhibitors to enzymes involved in sphingolipid metabolism, the researchers discovered that the drug-resistant leukemia cells were still sensitive to drugs that inhibited metabolic enzymes. These results, published in the Journal of Lipid Research in July, suggest that the metabolism of leukemias is a potential drug target for leukemias, particularly for some drug-resistant leukemias.
Mediating angiogenesis
Tumors can cause blood vessels to grow toward the tumor so that cancer cells can gain more nutrients, a process called angiogenesis. However, what chemical factors mediate this process are not fully understood. Rand et al. determined that, when the lipid EET is modified by the enzyme COX-2 to form modified EET, the modified EET promotes blood vessel formation. These results may explain the effectiveness of COX-2 inhibitors in slowing tumorigenesis and cancer development. The researchers published their study in JLR in October.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Glaucoma model links immune signaling to disease progression
Researchers at Duke University determine genetic variations that could increase the risk of developing glaucoma.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.

Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.

Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.

