From the journals: MCP
We offer summaries of recent papers in the journal Molecular & Cellular Proteomics.
A new role for an insulin receptor substrate
Insulin receptor substrate 2, or IRS2, is an adaptor molecule recruited by insulin receptors to facilitate insulin signaling. As such, IRS2 is implicated in diabetes mellitus and in breast and pancreatic cancers and neuroblastoma. IRS2 function is known to mediate signaling including lipid uptake and metabolism, and key checkpoints for its function are the ubiquitin ligases that can break down the proteins that help IRS2.
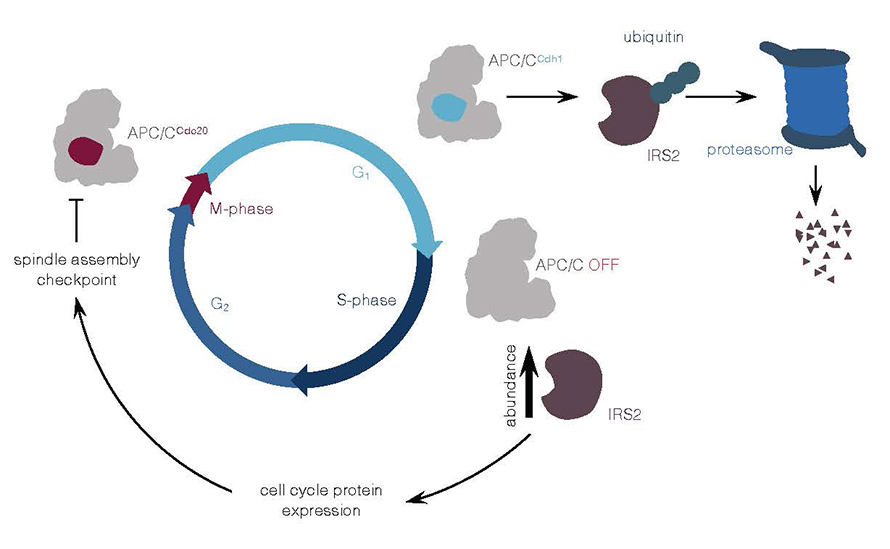
Though some researchers have reported ubiquitin ligases that target both IRS2 and other IRS molecules, none had reported ubiquitin ligases that specifically target IRS2. Sandhya Manohar and a team at Harvard Medical School addressed this question with a proteomic screen of anaphase-promoting complex/cyclosome, or APC/C, to identify potential substrates during the intermediate G1 growth phase of the cell cycle. The APC/C assists in the breakdown of proteins involved in the cell cycle, achieving this feat with the help of activators. In their recent paper in the journal Molecular & Cellular Proteomics, the authors provide evidence suggesting IRS2 is a direct substrate of APC/CCdh1, where Cdh1 is an APC/C co-activator during G1.
When APC/C was blocked and/or Cdh1 was knocked down, the researchers observed increased IRS2 levels. They also found that IRS2 depletion decreases the expression of cell-cycle regulatory proteins and perturbs the mitotic spindle checkpoint during the mitotic phase. The stabilization of IRS2 depends on a unique D-box motif present in IRS2, thereby establishing a new mode of regulating IRS2.
Together, these findings identify a ubiquitin ligase that targets only IRS2 and not the other main IRS protein, IRS1, and establish a new role for IRS2 as a cell-cycle checkpoint regulator. In addition to IRS2 roles in lipid uptake and metabolism, this study unravels a new role of IRS2 in cell-cycle regulation.
Creating a neural signaling roadmap
Neurotransmitters transmit chemical signals from synapses to facilitate cell–cell communication. Impairments in synapse function play a key role in the pathology of neurological diseases such as Parkinson’s, schizophrenia and epilepsy. Synaptosomes, which are stable and functional nerve terminals and can be manipulated for various processes, are employed to understand molecular processes of communication. These processes include post-translational modifications, or PTMs, which are important for protein function.
In a recent paper in the journal Molecular & Cellular Proteomics, Inga Boll and a team from the University of Southern Denmark write about the role of a PTM, sialylation (the addition or removal of sialic acid), in N-linked glycopeptides in rat synaptosomes. The researchers have made a large-scale comprehensive map of sialic acid–containing glycosylation sites. Protein kinases, proteases and phosphatases carry these sialylated glycosylation sites.
Brief depolarization of the synaptosomes allowed the researchers to identify changes in site-specific sialylated and desialylated N-linked glycoproteins. These changes occurred on essential synaptic proteins such as ion channels, transporters and synaptic vesicle proteins and are as dynamic as post-translational phosphorylation. This study unravels sialylation, a novel modulator that alters the neurotransmitter release in response to depolarization. The researchers have identified the need to establish state-of-the-art techniques to study the implications of the molecular functions of sialylation.
What ionizing radiation does to proteins
X-rays, gamma rays and ultraviolet rays are useful for disinfecting instruments and in diagnostic medical exams, but high doses of such radiation can damage tissues and lead to cancer. Cell death due to the production of reactive oxygen species, or ROS, such as hydrogen peroxide and superoxide can cause tissue damage. Researchers know that, at the DNA level, cells employ several strategies to avoid and repair DNA damage due to oxidation. However, the mechanisms involved in repairing the DNA damage are relatively unknown.
In a recent paper in the journal Molecular & Cellular Proteomics, Steven T. Bruckbauer and colleagues at the University of Wisconsin describe performing mass spectrometry on Escherichia colicells that they had treated with ionizing radiation to identify the resulting oxidative modifications. The researchers determined that fewer than 10% of the proteins present in the organism were affected by radiation. The most common oxidative modification was hydroxylation, followed by dioxidation. Glyceraldehyde 3′-phosphate dehydrogenase, or GAPDH, is the primary target of oxidation by radiation.
Similar oxidation on GAPDH is also present in human breast carcinoma cells, suggesting that it is a critical player in oxidative damage from bacteria to eukarya. This work adds to our knowledge of the repertoire of cellular functions of GAPDH.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.

