Stabilizing messenger RNA may lead to COVID-19 vaccine development
The whole world seems to be holding its collective breath for development of a vaccine against the novel coronavirus that causes COVID-19. Numerous groups are developing vaccines, and a few of their formulations are in clinical trials. Whereas traditional vaccine development involves inoculation with an attenuated form of the virus or with a noninfectious antigen derived from a viral protein, two groups, Moderna (in collaboration with the National Institutes of Health) and BioNTech (in collaboration with Pfizer) are taking a novel approach: directly administering to patients messenger RNA, or mRNA, that encodes viral antigens. Work from my lab carried out in collaboration with researchers in Poland and Germany over 30 years is playing a part in the BioNTech/Pfizer trials.
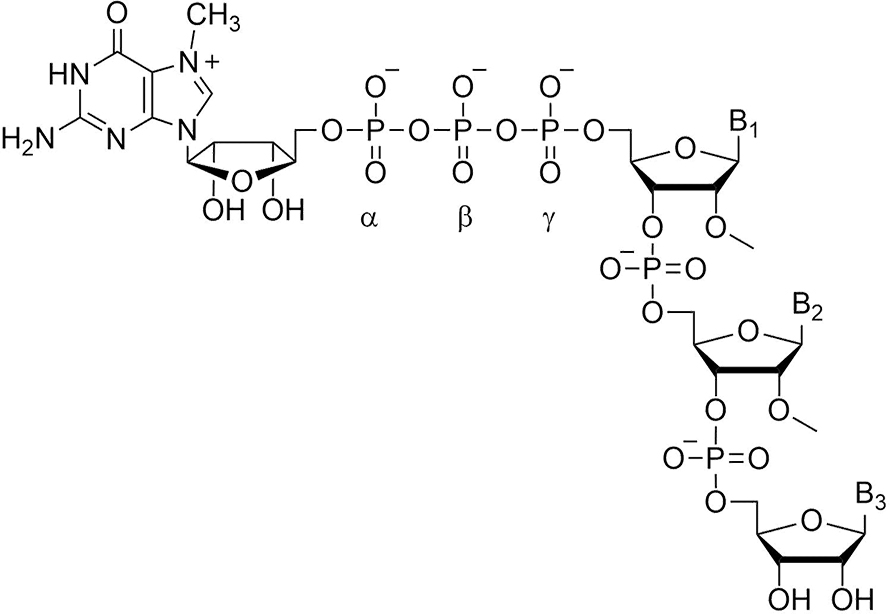
Our work centers around the cap of mRNA, a unique structure consisting of 7-methylguanosine linked 5′-to-5′ via a triphosphate bridge to the first encoded nucleotide of mRNA. I became interested in caps as a result of postdoctoral work in the lab of Bob Schimke, which I joined in 1970. Bob wanted to know whether cells producing very high proportions of a single protein did so because of gene amplification. Choosing the estrogen-stimulated chick oviduct as a model system, his strategy was to count genes by hybridization kinetics before and after estrogen administration. This was before the days of recombinant DNA, so we needed to purify ovalbumin mRNA by biochemical techniques, which in turn necessitated an assay for ovalbumin mRNA. The rabbit reticulocyte lysate system previously had been shown to synthesize duck hemoglobin in response to duck RNA. We found that oviduct RNA added to this system resulted in synthesis of ovalbumin; our paper and another one in in 1971 were the first to show mRNA-directed synthesis of a nonglobin protein in rabbit reticulocyte lysate.
At that time, mRNA was understood from genetic studies but virtually unexplored as a biochemical entity. The ability to obtain it in milligram quantities now opened up the possibility of a wide range of new studies. While still in the Schimke lab, I wrote my first NIH grant proposal after I had been offered an assistant professorship at University of Kentucky Medical Center. One of the specific aims was to determine the structure at the 5′-end of mRNA. I was unaware that several well-established labs already were engaged in this search. By 1975, before the first renewal of my NIH grant was due, Aaron Shatkin had scooped me completely. Fortunately, this was not a dead end — many fascinating things about mRNA remained to be explored.
The presence of a cap on mRNA greatly enhances its translational efficiency. Small-molecule analogs of the cap, such as mononucleotides and dinucleotides that include 7-methylguanosine, are useful tools to study initiation of protein synthesis because they compete with mRNA for recognition by the translational machinery. Early in my studies of the cap and initiation of translation, I began a long-lasting collaboration with Ed Darzynkiewicz of the University of Warsaw, who had done postdoctoral work with Aaron Shatkin to make cap analogs at about the same time I was starting my lab at the University of Kentucky. Ed and his colleagues synthesized a large collection of cap analog inhibitors of translation, but at one point we wished to synthesize entire mRNAs containing modified caps. A relatively simple method for this had been developed by Walter Fiers, but it led to cap dinucleotides being incorporated in both the normal and reversed orientation. We developed and patented what we called “anti-reverse cap analogs” by blocking the 3′-OH of 7-methylguanosine; these mRNAs had higher translational efficiency both in cell-free systems and when introduced into cultured cells.
The anti-reverse modification also made it possible to introduce structural changes at individual phosphate moieties that would be located unambiguously with respect to the body of the mRNA. We used this to explore whether changes in the phosphate moieties could alter the rate of mRNA degradation, an essential and highly regulated process for the control of gene expression. In one degradation pathway, the first step is cap removal by the pyrophosphatase complex Dcp1/Dcp2, and then the RNA chain is hydrolyzed in the 5′-to-3′ (front to back) direction by the exonuclease Xrn1. Pyrophosphatase cleavage occurs between the alpha and beta phosphate moieties.
Ewa Grudzien–Nogalska, a postdoc in my lab at the Louisiana State University Health Sciences Center and previously a Ph.D. student in Ed’s lab at the University of Warsaw, examined cap analogs that, when incorporated into mRNA, retarded mRNA degradation. Collaborating with one of Ed’s colleagues, Jacek Jemielty, Ewa found that substituting a methylene group for the alpha-beta bridging oxygen prevented Dcp1/Dcp2 from cleaving the triphosphate bridge, which in turn stabilized the mRNA against 5′-to-3′ degradation in cultured cells. Unfortunately, this modification also reduced the translational efficiency of the mRNA.
Next, Joanna Kowalska, a Ph.D. student in Jacek’s lab, synthesized cap dinucleotides in which sulfur was substituted separately for each of the nonbridging oxygen atoms in the triphosphate chain. Two isomers of the sulfur-for-oxygen substitution at the beta phosphate were resistant to Dcp1/Dcp2 hydrolysis and, when incorporated into mRNA, conferred enhanced stability in cultured cells. Surprisingly, they were translated more efficiently than mRNAs capped by the parent (oxygen-containing) dinucleotide. This technology was patented by LSU and University of Warsaw. Another series in which nonbridging oxygens were substituted with BH4 groups was synthesized, patented and found to have the same property of both higher translational efficiency and higher stability.
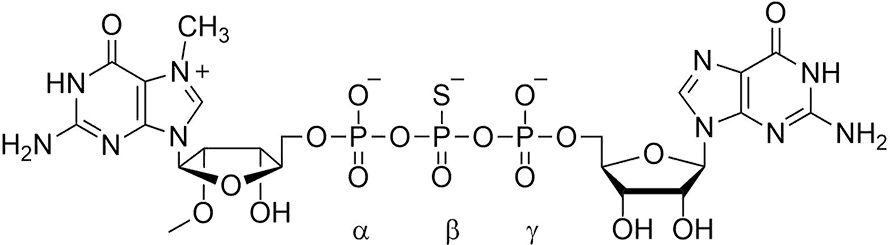
The ability to make synthetic mRNA that was more stable in cultured cells attracted the attention of Ugur Sahin, an oncologist at Johannes Gutenberg University in Mainz, Germany, who had long sought personalized vaccines for cancer immunotherapy. When Ed and I attended the RNA Society meeting in Berlin in 2008, Ugur drove up from Mainz to have dinner with us and explore the idea of using our stabilizing cap analogs in the clinic. The plan he described, and ultimately accomplished, was to sequence all the RNA from a patient’s tumor to discover all proteins being expressed, compare them to the canonical proteins to find aberrant proteins synthesized by the cancer cells, make synthetic mRNA encoding these neoepitopes, and then inject it into patients where it would be taken up and translated by antigen-presenting cells, thereby activating T cells that would attack the patient’s cancer cells. His plan was astonishing to me because mRNAs are unstable in the lab and cell and most mammalian cells are extremely inefficient in taking up intact nucleic acids. However, I subsequently learned that a type of antigen-presenting cell, the immature dendritic cell, is different from most cells because it takes up mRNA by micropinocytosis, being specialized in sampling its environment for potential pathogens. Ugur’s group was interested in our mRNA-stabilizing cap structures because mRNA persistence in dendritic cells results in more translation, more neoepitopes produced, and a greater T cell response. They subsequently demonstrated that this occurred with a model antigen in mice.
To realize the goal of RNA-based cancer immunotherapy, Ugur founded BioNTech in Mainz in 2008, which then licensed the patents for our mRNA-stabilizing cap analogs from LSU and the University of Warsaw. Clinical trials for safety with malignant melanoma patients produced favorable results. BioNTech then partnered with Genentech to pursue RNA-based cancer immunotherapy trials for efficacy and comparison to standard treatments.
When the novel coronavirus appeared In China last year, BioNTech already was experienced in using mRNA to produce an immunological response, so it moved quickly to partner with Pfizer to develop a COVID-19 vaccine. German clinical trials began in April and U.S. clinical trials began in May. Four formulations are being tested, two of which utilize cleavage-resistant caps and two of which utilize other features that enhance translational yield. Compared with traditional vaccine development approaches, which may require 12 to 18 months, production of mRNA-based vaccines can be as short as four months.
Yogi Berra is credited with observing, “It’s tough to make predictions, especially about the future.” We don’t know which vaccine approach will be the most successful. However, it is clear that only when a vaccine is available will we have protection against COVID-19 and its destruction of health and economic well-being. A silver lining to this dark cloud is that we likely will have new tools with which to develop vaccines more rapidly against the next dread disease that appears, something that likely will be the rule rather than the exception in our increasingly interconnected world. The speed and flexibility of inducing an mRNA-based immunological response makes it likely that this will be a major component of our toolbox.
Disclosure
Louisiana State University is one owner of the patents for mRNA-stabilizing cap analogs that are currently licensed by BioNTech. As one of the inventors, Robert Rhoads receives from LSU a small percentage of income generated by these patents. To date, no income has been derived from the COVID-19 initiative.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
Innovative platform empowers scientists to transform venoms into therapeutics
Scientists combine phage display and a “metavenome” library to discover new drugs that bind clinically relevant human cell receptors. Read about this recent Molecular & Cellular Proteomics paper.

Meet Shannon Reilly
The JLR junior associate editor discusses the role of adipocytes in obesity at Weill Cornell Medical School.

Meet Donita Brady
Donita Brady is an associate professor of cancer biology and an associate editor of the Journal of Biological Chemistry, who studies metalloallostery in cancer.

Glyco get-together exploring health and disease
Meet the co-chairs of the 2025 ASBMB meeting on O-GlcNAcylation to be held July 10–13, 2025, in Durham, North Carolina. Learn about the latest in the field and meet families affected by diseases associated with this pathway.
Targeting toxins to treat whooping cough
Scientists find that liver protein inhibits of pertussis toxin, offering a potential new treatment for bacterial respiratory disease. Read more about this recent study from the Journal of Biological Chemistry.

Elusive zebrafish enzyme in lipid secretion
Scientists discover that triacylglycerol synthesis enzyme drives lipoproteins secretion rather than lipid droplet storage. Read more about this recent study from the Journal of Biological Chemistry.

