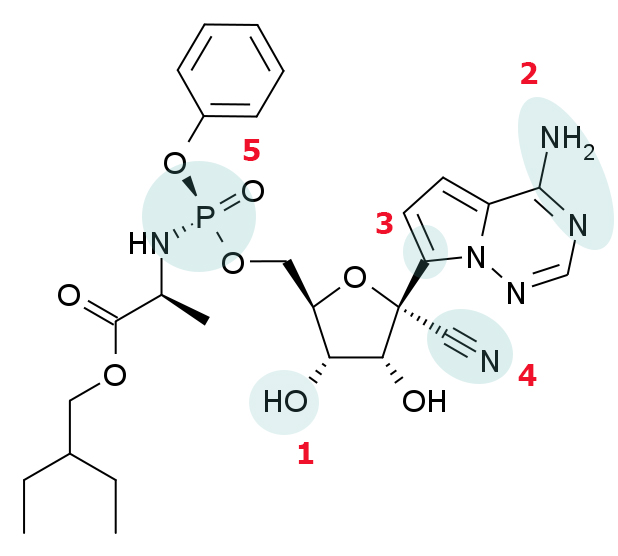
Anatomy of a molecule: What makes remdesivir unique?
The World Health Organization in late January convened experts to discuss experimental therapeutics for patients with the emerging coronavirus with no name, no vaccine and no treatment. The panel reported that “among the different therapeutic options, remdesivir was considered the most promising candidate.”
Within weeks, a clinical trial of the compound was underway in China. Results are expected in April; in the meantime, the outbreak of SARS-nCoV-2, the virus that causes COVID-19, has become a global pandemic.
Remdesivir is a nucleoside analog, one of the oldest classes of antiviral drugs. It works by blocking the RNA polymerase that coronaviruses and related RNA viruses need to replicate their genomes and proliferate in the host body.
The molecule originally was synthesized as part of a screen for inhibitors of the hepatitis C virus RNA polymerase. Its inventors at Gilead Sciences decided to move forward with a different nucleoside analog compound to treat hepatitis C. But RNA-dependent RNA polymerases are conserved between many viruses. Experiments in vitro, in cell culture and in animal models have shown that remdesivir has broad-spectrum activity against RNA viruses, including filoviruses (like the one that causes Ebola) and coronaviruses.
Remdesivir resembles the RNA base adenosine, shown here as a monophosphate.
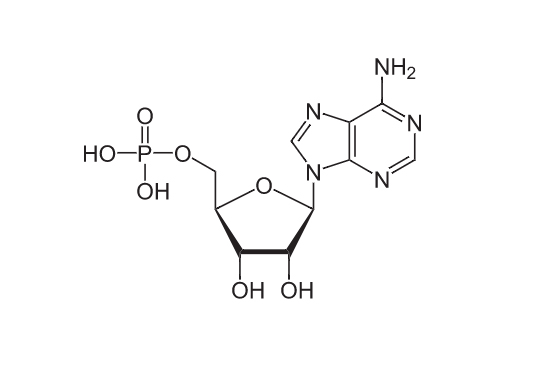
The compound and ATP have some important differences, but some features are very similar. ASBMB Today spoke to medicinal chemist Katherine Seley–Radtke at the University of Maryland, Baltimore County, and structural virologist Craig Cameron at the University of North Carolina, Chapel Hill about what makes the molecule interesting. Click on a feature marked in blue to read their remarks.
3’ hydroxy group
Different classes of nucleoside/nucleotide analogs have different effects on polymerases. Remdesivir is in a class called nonobligate chain terminators, because it should, in theory, be possible to add more nucleotides to a strand of RNA after remdesivir has been added due to the presence of the hydroxyl group at carbon 3 in the sugar.
“That hydroxy group is what is required for continued synthesis of nucleic acid, whether it be RNA or DNA,” said virologist Craig Cameron, a professor at the University of North Carolina at Chapel Hill who studies the interactions between nucleoside analogs and viral polymerases.
Recent research suggests that when mixed with RNA polymerases from coronaviruses or flaviviruses in vitro, remdesivir doesn’t terminate the synthesis of a new RNA strand right away. Instead, Cameron said, “It takes a few cycles of nucleotide addition before you can see the termination effect.”
Those additional nucleotides may help shield remdesivir from coronavirus proofreading enzymes that are known to remove unnatural nucleotide analogs.
Base pairing to uracil
In adenosine in double-stranded RNA, this face of the molecule is involved in base-pairing with uracil. The two nitrogens act as proton donor and acceptor, respectively for hydrogen bonds to atoms in the uracil base.
Chemists think that remdesivir, by presenting a very similar binding face, gets incorporated into a growing RNA strand by viral polymerases.
C-nucleoside bond
The link between ribose and the base is called the glycosidic bond. Usually, it connects the 1’ carbon in the ribose ring to a nitrogen in the base. But in remdesivir (and some other nucleotide analogs) the sugar and the nucleobase are connected by a bond between two carbons.
“It definitely provides much greater stability (against) nucleases and other enzymes that can cleave the nucleobase from the sugar,” said Katherine Seley-Radtke, a medicinal chemist at the University of Maryland, Baltimore County who works on the design and synthesis of antiviral nucleoside analogues. With a C-nucleoside, “you’d have to break a carbon-carbon bond, whereas in a normal nucleoside you’re breaking a hemi-aminal bond, which is actually fairly unstable. So having that carbon-carbon bond is a great advantage.”
1’ cyano group
Ask a group of chemists what jumps out at them about remdesivir, and most will start with this dramatic feature. Substitution at this carbon is unusual, and probably only possible because of the strength of the C-nucleoside bond.
According to an article in the Journal of Medicinal Chemistry, the cyano group was initially added because a precursor molecule, a very effective inhibitor of viral RNA polymerases, also blocked the mitochondrial RNA polymerase in mice. To make a molecule without those toxic side effects, chemists at Gilead tried a series of substitutions at the 1’ carbon. The compound with the cyano group worked best: it still blocked the hepatitis C polymerase, but was no longer incorporated by host cell polymerases.
“You can’t predict activity. You have to make it and test it,” Seley-Radtke said. “But even small changes can have amazing consequences.”
Phosphate
“You see all that flotsam and jetsam coming off at the 5’ hydroxyl?” said Katherine Seley-Radtke. Among medicinal chemists, this type of protecting group is casually known as “a McGuigan protide.” Designed by medicinal chemist Chris McGuigan in the 1990s, this type of protecting group and its variations are widely used to deliver nucleotide analogs into cells.
“It is a brilliant system, because it accomplishes two things,” Seley-Radtke said. “No. 1, an issue with nucleosides is that they’re polar and their phosphates are even more polar.” Masking the highly negative phosphate groups with esters or amides reduces the molecule’s overall polarity, letting it cross the plasma membrane into cells.
Second, in order to be recognized by polymerases, the analog needs to resemble a normal nucleotide triphosphate—which means it needs to be phosphorylated.
“The first phosphorylation, either by cellular or viral kinases, is oftentimes very difficult,” Seley-Radtke said. “A lot of those kinases are very, very picky in terms of recognition.” By arriving in the cell with its first phosphate already in tow, remdesivir and related nucleotide analogs skip that rate-limiting step. After the protecting groups are cleaved, the nucleotide analog is a reasonable substrate for later nucletodie kinases.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
Bacterial enzyme catalyzes body odor compound formation
Researchers identify a skin-resident Staphylococcus hominis dipeptidase involved in creating sulfur-containing secretions. Read more about this recent Journal of Biological Chemistry paper.

Neurobiology of stress and substance use
MOSAIC scholar and proud Latino, Bryan Cruz of Scripps Research Institute studies the neurochemical origins of PTSD-related alcohol use using a multidisciplinary approach.
Pesticide disrupts neuronal potentiation
New research reveals how deltamethrin may disrupt brain development by altering the protein cargo of brain-derived extracellular vesicles. Read more about this recent Molecular & Cellular Proteomics article.

A look into the rice glycoproteome
Researchers mapped posttranslational modifications in Oryza sativa, revealing hundreds of alterations tied to key plant processes. Read more about this recent Molecular & Cellular Proteomics paper.
Proteomic variation in heart tissues
By tracking protein changes in stem cell–derived heart cells, researchers from Cedars-Sinai uncovered surprising diversity — including a potential new cell type — that could reshape how we study and treat heart disease.

Parsing plant pigment pathways
Erich Grotewold of Michigan State University, an ASBMB Breakthroughs speaker, discusses his work on the genetic regulation of flavonoid biosynthesis.