The cellular conundrums of hepatitis C
Hepatitis C is a rarity among the viruses that infect humans on a global scale. Although an effective vaccine still is being developed for it, properly administered antivirals can eliminate the virus in 95% of the people who are infected with it.

Alter and Michael Houghton, also won the Lasker Award in 2016 with Ralf
Bartenshlager and Michael Sofia for developing a system to study the replication
of the virus that causes hepatitis C.
But it hasn't been an easy nut to crack. When the virus was isolated in 1989, scientists ran into repeated dead ends as they tried to get it to replicate in cell culture. The journey to develop an effective system in which to grow and study HCV involved hundreds of scientists and took more than 15 years.
In October, Charles Rice at The Rockefeller University, along with Harvey J. Alter at the National Institutes of Health and Michael Houghton at the University of Alberta, won the Nobel Prize in physiology or medicine for their seminal discoveries leading to the identification of the hepatitis C virus.
"It was a huge surprise to me, and a shock, and a very humbling experience," said Rice. "When you think about the many people that have been involved in this success story, it's something that has spanned decades and involved thousands of dedicated scientists and clinicians to really get to where we are today."
Epidemic jaundice
The Hippocratic physicians are credited with the first descriptions of contagious jaundice in the fifth century B.C. The condition, the hallmarks of which are yellowing of the skin and the whites of the eyes and dark urine, occurs when liver inflammation or a blocked bile duct allows bilirubin to build up in the blood.
"The connection made between the liver and jaundice was remarkable, bearing in mind that the Hippocratic physicians had not performed dissections and that their medical views were based on observation," Niki Papavramidou, Elizabeth Fee and Helen Christopoulou–Aletra wrote in the Journal of Gastrointestinal Surgery in 2007.
Epidemic jaundice was recorded in the 17th and 18th centuries. Tens of thousands of Union soldiers during the American Civil War were sickened.
By the 1940s, scientists had begun to differentiate between hepatitis A and B by comparing their incubation times.
During World War II, hundreds of thousands of American and European troops contracted viral hepatitis. In one case, upward of 330,000 U.S. service members contracted HBV after being given a yellow fever vaccination that contained contaminated human blood serum.
Medical anthropologist Baruch "Barry" Blumberg discovered HBV serendipitously in 1967. He was interested in how inherited traits make people more or less susceptible to disease and for his studies traveled the globe collecting blood samples from remote populations.
He found a surface antigen for hepatitis B in the blood of an Australian aborigine, which led to the identification of the virus and its role in causing acute and chronic hepatitis and liver cancer.
In 1976, Blumberg, along with D. Carleton Gajdusek, won the Nobel Prize for physiology or medicine for their work on the virus. Blumberg's birthday, July 28, was adopted as World Hepatitis Day in 2008.
Not A or B, but C
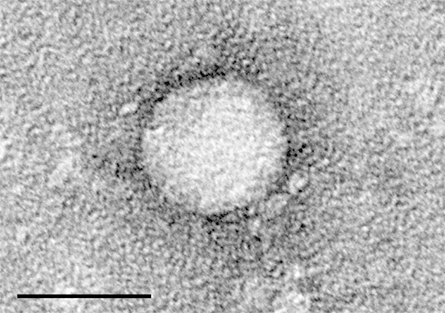
infects approximately 71 million people worldwide.
In the early 1970s, National Institutes of Health physician Harvey J. Alter was investigating hepatitis in patients who had received blood transfusions but had tested negative for both the A and B viruses.
The new illness, then known as "non-A, non-B" hepatitis and now known as hepatitis C, became a priority for Michael Houghton, who was working for the pharmaceutical firm Chiron at the time.
In 1989, Houghton and his colleagues Qui-Lim Choo and George Kuo isolated a near-complete genetic sequence of HCV, which allowed other scientists to develop blood-based screening tests for it.
That same year, Charles Rice, then at the Washington University School of Medicine in St. Louis, developed the first infectious clone of yellow fever virus, the prototype flavivirus in the family Flaviviridae, which also includes the mosquito-borne dengue, Zika and West Nile viruses.
Not long after, Rice received a call from Stephen Feinstone at the U.S. Food and Drug Administration, who wanted him to retool his work to tackle this new flaviviruslike agent — HCV.
Rice's flaviviral investigations
After graduating with a bachelor's degree in zoology from the University of California, Davis, in 1974, Rice began graduate school at the California Institute of Technology. He initially planned to continue his undergraduate research in developmental biology, but his first lab placement was with virologist James H. Strauss Jr.
"I started working in the Strauss lab, which was really the only active virology lab at Caltech," Rice said. "And I really enjoyed that, so I just stayed there."
Near the end of his doctoral work, completed in 1981, Rice began studying the mosquito-borne yellow fever virus, after which both the genus Flavivirus and family Flaviviridae are named. He continued his research as a postdoctoral fellow in Strauss' lab for the next four years.
After briefly working at the Australian National University with Lynn Dalgarno, who had co-discovered with John Shine in 1973 the Shine–Dalgarno sequence involved in mRNA translation initiation, Rice took a faculty position at WUSTL in 1986. There, he continued to work with yellow fever virus.
"At Caltech, we'd already determined the genome structure of the virus and had some ideas about what the expression strategy might be for the viral proteins," Rice said. "But at Wash U, I was actually trying to build an infectious clone for yellow fever that would allow us to do more directed genetic manipulations on this virus to understand more about how it works."
When Rice was setting up his lab in early 1987, the first person he hired was Arash Grakoui, who had just earned an undergraduate degree and was looking for employment as a lab tech.

was starting his lab at WUSTL, has gone on several excursions to Wyoming
over the years with Rice (right), whom he describes as an avid outdoorsman.
"It was Charlie and me, and we had a bench, sitting across from one another. And soon there was a graduate student, and then a postdoc as well," said Grakoui, now a professor of medicine at Emory University. "Right off the bat, he trained me as a student or postdoc in the lab and let me have a lot of freedom to be creative."
The next few years also saw a spate of more unusual hires, and Rice's lab began working with HCV. Brett Lindenbach, a professor of microbiology at Yale University, was a graduate student in Rice's lab through the 1990s and worked closely with Alexander "Sasha" Kolykhalov.
"After the Soviet Union collapsed, a lot of very excellent — like, literally, some of the top Russian virologists — just came and worked in Charlie's lab more or less as postdocs," said Lindenbach, who also did his second postdoctoral fellowship with Rice at The Rockefeller University. "Most of them came from Novosibirsk, which is where the Soviet Union did its bioweapons virus research."
Kolykhalov and Rice began building consensus clones, full-length complementary DNAs (called cDNAs for short) of HCV that are cloned within bacterial plasmids, but the clones were never functional in cell culture or in chimpanzees, the only other animal the virus is known to infect, Lindenbach said. This led them to believe that they might be dealing with an incomplete genome.
"Charlie and Sasha set out to try and discover what the real 5' and 3' ends of the viral genome were," Lindenbach said. "They confirmed that the 5' end was indeed correct, but then they discovered this big piece of the 3' end that had been missing."
Kolykhalov, Rice and Feinstone reported their findings about a highly conserved sequence element at the 3' end of the HCV genome in 1996 in the Journal of Virology. The next year, Rice reconstructed a full-length consensus clone, a single cDNA clone representing the average of variants within the viral population, but still had no system to test it in.
Despite the researchers' best efforts, the variant refused to replicate in any established cell lines. So Feinstone injected the viral variant into the livers of two chimps. (Author's note: In 2015, the National Institutes of Health announced that it would no longer support biomedical research involving chimpanzees.)
"This was a Hail Mary sort of experiment, because this is a 10,000-base-pair-long RNA," Rice said. "And if you're just injecting it into an organ in vivo and hoping that somehow it gets inside of a cell so it can be translated by that machinery, you're hoping to get lucky."
Rice, Feinstone and their co-authors did get lucky: The virus caused changes in blood and pathology resembling those seen in humans with hepatitis C, providing direct evidence that HCV was responsible for the transfusion-mediated hepatitis.
Their 1997 paper, published in the journal Science, noted that the scientists had fulfilled Koch's postulates on a molecular level: An infectious organism must be present during each case of a disease and must able to be isolated and purified, and the organism must cause the disease when a new host is infected with it.
"I recall pointing out that this helps to prove that HCV is the causative agent of hepatitis C, because it fulfills sort of a molecular form of Koch's postulates," Lindenbach said. "And Charlie thought that was a great idea and wrote that into the paper."
Hepatic breakthrough
While the researchers lacked a way to study the virus in cell culture, a model arrived in 1999. Ralf Bartenschlager, a professor at the University of Mainz, and Volker Lohmann figured out how to select for a variant of the virus to replicate in HuH7 cells originally derived from a cancerous tumor in human liver through use of a subgenomic replicon system.

educational pamphlets about testing and diagnoses for hepatitis C
virus, with which nearly 4 million Americans were then infected.
"What they did was to chop out a chunk of the genome that encoded what we believed to be the structural proteins of the virus, the glycoproteins and the capsid protein, and replace that with a drug-selectable marker," Rice said.
That marker was a neomycin gene that confers resistance to the drug G418, to which human cells are sensitive. By exposing the hepatic cells to G418, the cells that did not support the HCV replicon and take up the drug resistance gene were wiped out, leaving only HCV-laden cells for the researchers to pore over.
"If you're looking for a needle in a haystack, you need a big magnet. And that's what this was," Lindenbach said.
Rice and his colleagues then found that variants of HCV in the subgenomic replicon system harbored mutations that were adaptive to their environment. When those mutations then were engineered into new replicons, the researchers found that they could increase the efficiency of the system more than 10,000-fold.
Another advance came in 2002 when Keril Blight, one of Rice's postdocs, helped cure cells of replicons to see if there might be a selection for more HCV replicon-permissive cellular environments. The increased permissiveness of one of the cured cell lines allowed the researchers to detect HCV RNA and antigens early after RNA transfection, which eliminated the need for selection of replication-positive cells.
The final breakthroughs for growing hepatitis C arrived in 2005, when a team that included Japanese virologist Takaji Wakita and Bartenschlager found that a rare isolate of HCV from a patient in Japan with acute fulminant disease was able to undergo its entire life cycle in cell culture without induced mutations. At the same time, Rice's group, including Lindenbach, described a chimeric full-length HCV genome that replicated and produced virus particles that were infectious in cell culture, and a group led by Frank Chisari at Scripps Research reported a robust HCV replication system that used the JFH-1 molecular clone and HuH-7-derived cell lines.
In the years that followed, researchers were able to develop direct-acting antivirals for hepatitis C. Until that point, the viral infection had been treated with the more broadly acting antivirals pegylated interferon and ribavirin, a cocktail that carried significant side effects and only boasted a 50% success rate of eliminating the virus, Rice said.
In 2014, the FDA approved the direct-acting combination pill Harvoni made up of the drugs ledipasvir and sofosbuvir, which are effective in eliminating HCV in more than 95% of the people who take them.
According to Craig Cameron, a virologist at the University of North Carolina at Chapel Hill, while HCV is able to develop resistance to individual drugs, a combination of antivirals that target different viral functions kills the virus outright.
"If you hit it hard enough, long enough, you can force it to extinction," Cameron said.
Nice catch
In 2001, Rice moved his lab to Rockefeller in New York. In 2002, he convinced Grakoui, who had enrolled as a graduate student at WUSTL in the lab of immunologist Paul Allen in 1994 after seven years in Rice's lab, to join him as a postdoctoral fellow.
"Coming back as a postdoc really enabled me to bring along all the tools that I had gathered in graduate school to tackle some of the questions that we were interested in about viral immunology," he said.
During their years at WUSTL, Rice, an avid outdoorsman, had introduced Grakoui to fishing through long excursions to Wyoming.
"Charlie described to me on one of our trips that fishing is a lot like science: You don't actually expect to catch anything, but if you do, it is incredible," Grakoui said. "You have done the best you can, and you just let nature takes its course."

The ABCDEs of hepatitis
While overconsumption of alcohol, environmental toxins and autoimmune disease play significant roles in global incidence of liver inflammation, or hepatitis, viral infections cause the majority of cases.
The Hippocratic physicians in the 5th century B.C. identified five types of jaundice based upon the color of patients' skin, urine and feces and other symptoms. By coincidence, there are five hepatitis-causing viruses.
Hepatitis A, B, C and E viruses all cause liver inflammation, but none of the viruses shares close lineage.
Hepatitis A and E are transmitted by contaminated food or water and cause acute disease.
Hepatitis B and C, as well as hepatitis D, which can propagate only in the presence of hepatitis B, are chronic blood-borne pathogens that spread through intravenous drug use and blood transfusions.
Together, blood-borne hepatitis viruses, which result in cirrhosis and liver cancer, cause more than 1 million deaths each year.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in People
People highlights or most popular articles

2026 PROLAB awardees announced
Each of the seven awardees will work in labs across Canada and the United States and will receive $7,000 to fund travel and living expenses.

Uncovering the molecular roots of fatty liver disease
Physician–scientist Silvia Sookoian discusses her path from hepatitis C care to MASLD research, her use of multi-omics to study steatotic liver disease, and how lipid metabolism and genetics are reshaping understanding of MASH and liver health.

Kimble honored for lifetime achievement in genetics
She received the 2026 Thomas Hunt Morgan Medal and will be honored with a dedicated online profile and seminar.

Janetka named distinguished professor
Washington University awarded him the inaugural Carl Frieden Distinguished Professorship.

ASBMB members receive ASPET awards
The American Society for Pharmacology and Experimental Therapeutics awards Simone Brixius–Anderko, Paul Insel, Sudarshan Rajagopal, Emily Scott, Alan Smrcka and Jürgen Wess for their excellent research and mentoring work in pharmacology.

Kozul honored by Washington University
She received the 2025 Elliot L. Elson Education and Training Award.

