JLR: A close-up of nascent HDL formation
Oil and water don’t mix. But our aqueous blood is full of hydrophobic lipids — including cholesterol. To travel via the bloodstream, those lipids must hitch a ride on an amphipathic carrier. In a paper in the Journal of Lipid Research, scientists at Boston University report an advance in our mechanistic understanding of how one such carrier forms.
“Lipoproteins are like boats that deliver and remove cargoes of fatty substances to and from our cells,” said David Atkinson, chair of the physiology and biophysics department at Boston University School of Medicine and senior author on the JLR paper.
The subset of those “boats” that carry cholesterol and other lipids to the liver from other parts of the body are called high-density lipoproteins, or HDL, aka “good cholesterol.” HDL can remove cholesterol from distal cells — such as macrophages in the walls of arteries, where cholesterol accumulation can lead to heart attacks — and deliver it to liver cells, a process known as reverse cholesterol transport. The liver disposes of excess cholesterol by converting it into bile acids secreted into the small intestine.
According to Atkinson, a biophysicist, most of what is known about HDL formation comes from experiments that take a cell biological tack. In such studies, he said, “You can see (HDL formation) happening, and you can quantitate what happens, but you don’t understand the driving interactions that cause it to happen.”
HDL is built on a scaffold protein, apolipoprotein A-I. This apoA-I is thought to collect cholesterol and phospholipids from the cell membrane. Atkinson’s team wanted to better understand that process.
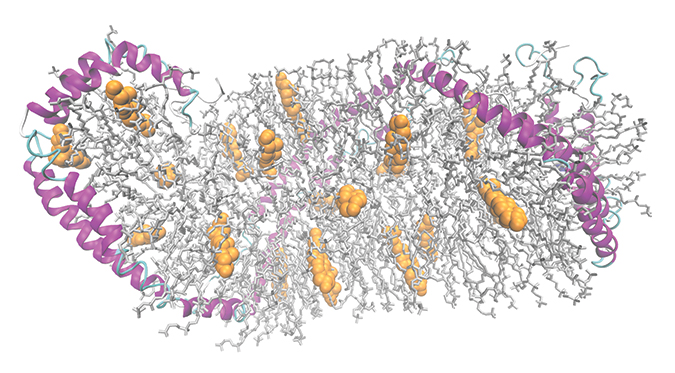 A model of a high-density lipoprotein particle shows apolipoprotein A-I in pink, phospholipids in gray and cholesterol in yellow.Wu et al/JBC 2009ApoA-I depends on a lipid transporter protein, ABCA1, that pumps cholesterol from the inner to the outer leaflet of the cell membrane. Because the cholesterol that ABCA1 transfers usually ends up bound to apoA-I, some researchers suspected a physical interaction between apoA-I and ABCA1. Others argued that cholesterol and phospholipids could diffuse passively and bind to apoA-I.
A model of a high-density lipoprotein particle shows apolipoprotein A-I in pink, phospholipids in gray and cholesterol in yellow.Wu et al/JBC 2009ApoA-I depends on a lipid transporter protein, ABCA1, that pumps cholesterol from the inner to the outer leaflet of the cell membrane. Because the cholesterol that ABCA1 transfers usually ends up bound to apoA-I, some researchers suspected a physical interaction between apoA-I and ABCA1. Others argued that cholesterol and phospholipids could diffuse passively and bind to apoA-I.
“Even if you demonstrate that apoA-I binds to the cell surface, you don’t actually know that it’s bound to ABCA1. It’s just bound to the cell surface,” Atkinson said. So he asked his team to see if they could “demonstrate that interaction actually happening in the isolated components.”
The team, led by graduate student Minjing Liu and supported by Xiaohu Mei and Haya Herscovitz, used isolated apoA-I and ABCA1 to test for a physical interaction. They were able to show immunoprecipitation of apoA-I with purified ABCA1.
The lab earlier had designed a mutant apoA-I with a little extra wiggle in an already flexible hinge region. For this study, they used the mutant to show that higher flexibility increased apoA-I lipidation, or the formation of nascent HDL. The team has not yet tested whether the extra-flexible mutant binds to ABCA1 better or whether binding of either form of apoA-I activates ABCA1.
But about one thing Atkinson is certain: “It’s the apoA-I/ABCA1 interaction which then enables the nascent HDL particle formation to happen as the membrane components are being transported out by ABCA1.”
Increasing reverse cholesterol transport may be a way to reduce atherosclerosis and heart disease. Atkinson is optimistic about the promise of understanding the physiological processes better.
“Translational research might be in vogue,” he said, “but remember that if you don’t do foundational basic discovery research, you will not have anything to translate.”
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.

