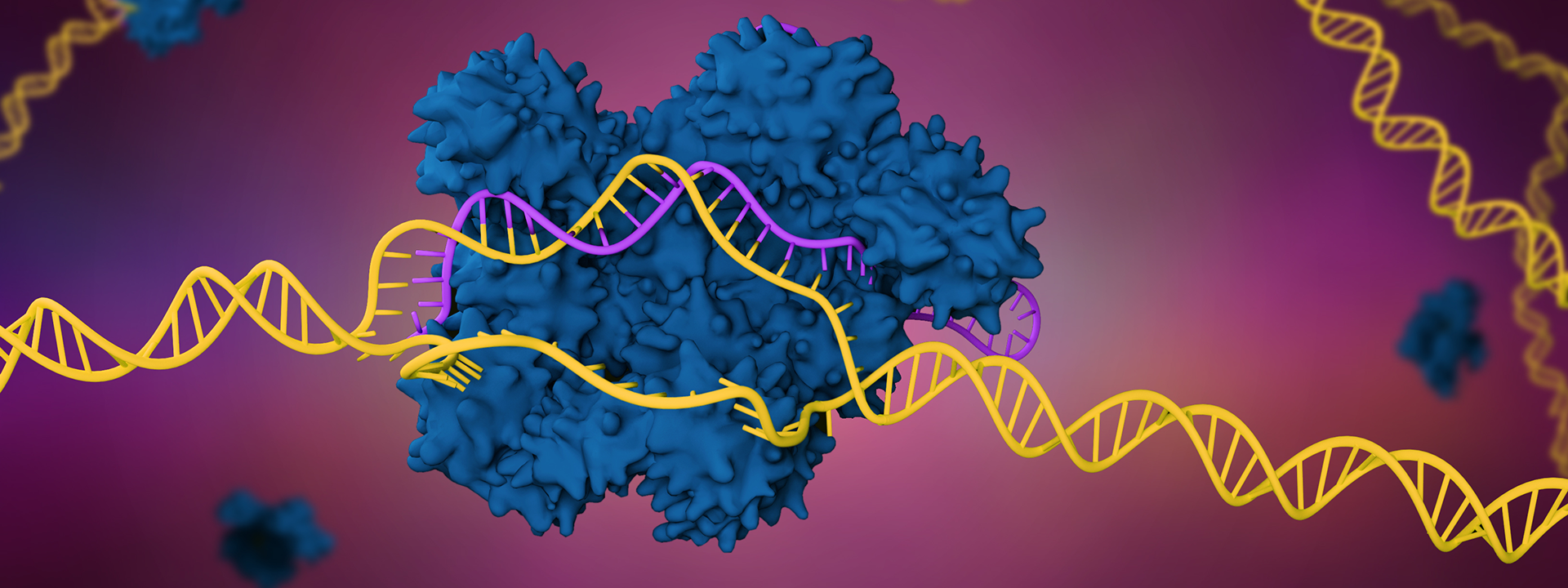
From bacterial immunity to scientific revolution
What can I tell you about CRISPR that you haven't heard before?
The technology has been the subject of countless journal clubs and chalk talks; it has exploded in citation count and triggered a wave of creative applications that has yet to crest. It has swept through biology labs, enabling researchers to modify genomes from Aspergillus to zebra mussel.
No matter how groundbreaking a discovery is, by the time it wins the Nobel Prize, the dust usually has settled. The finding or technique matures, becoming another slab in the foundations of science.
CRISPR is different. Some postdocs now using the technology as a matter of course were graduate students when its potential first was grasped. The most important molecular biology discovery of the 21st century is only 8 years old.

of San Francisco's Innovative Genomic Institute.
A strange season to celebrate
On the day it was announced that this year's Nobel Prize in chemistry was awarded to Emmanuelle Charpentier and Jennifer Doudna for their joint contribution to the use of CRISPR–Cas9 as a technology for genome editing, Charpentier remarked ruefully to an interviewer, "You connect to (the news), but you think it's another person, or it's surreal. … Now it's real, and now I have to deal with it."
A month later, Doudna told this reporter, "It's been a whirlwind, as you can imagine. But there's so much going on in the world right now that's a bit horrifying, but also motivating for scientists."
Before the Nobel announcement on Oct. 7, Doudna was focused on pandemic response at the Innovative Genomics Institute, which she founded in 2014 and co-leads. That work includes both a clinical testing program launched in the spring (see ASBMB Today coverage), and a more recently announced diagnostic technique for using CRISPR ribonucleoproteins and a cell phone to detect SARS-CoV-2 in patient samples (see CRISPR vs. COVID-19).

for the Science of Pathogens in and a professor at the Humboldt University
in Berlin.
"I wish we weren't having to do that, obviously," Doudna said of the IGI's all-hands-on-deck pandemic response. Then she added, "I think this is what science needs to do. As academics, we're not always trained to do this. But … we need to be nimble, and we need to be able to ask ourselves: How does our expertise apply in this moment?"
Megan Hochstrasser earned her Ph.D. in the Doudna lab working on CRISPR and now does outreach and education at the IGI, which she views as Doudna's effort "to make sure CRISPR leaves the lab and reaches the real world — and on top of that, make sure that the way it reaches the world is positive."
Hochstrasser has witnessed the evolution of CRISPR from when she was a grad student, when she got "to see a technology develop into something wildly important, when it started out as kind of an esoteric thing." Since moving to the IGI, she has watched it "grow from being very tiny to starting to get much bigger and more impactful."

It also brings together a team of scientists who can pool their expertise. In many cases, Doudna said, "There's no one of us that has the whole package of expertise … so we need to work together to do something that no one of us would be capable of doing on our own."
Collaboration is exactly how Doudna got into the CRISPR field in the first place.
A fashionable topic
Nobel-winning duos in the biochemical sciences often are scientists who have spent a significant fraction of their careers working together. Doudna and Charpentier's collaboration, in contrast, was brief and electrifying. They published just two research articles together — two articles with massive impact.
The idea of programmable genome engineering began long before their CRISPR work did. Restriction enzymes, which cut DNA at specific sequences, first were described in the 1960s. The discovery led researchers to reason that if they could alter a restriction enzyme's cut site, they could modify DNA sequences with exquisite precision. For a time in the early 2000s, researchers interested in programmable DNA cutting focused on two classes of proteins called zinc finger nucleases and transcription activatorlike effector nucleases, or TALENs. Both are made up of modular DNA-binding protein domains, which can be daisy chained to recognize a given sequence. But both proved difficult to engineer.
Doudna and Charpentier demonstrated that a new programmable system for DNA cutting could be simplified to just two components: one protein and one RNA.
Microbiologists had been aware for some time of a curious repeating feature in bacterial genomes called clustered regularly interspaced short palindromic repeats, or CRISPR for short (see Yogurt research, broad applications). In 2005, Spanish scientists discovered that the spacer sequences between the repeats matched up to the genes of viruses that infect bacteria. They proposed that the system might represent a kind of bacterial adaptive immunity: a way to recollect and respond to pathogens encountered before.

"CRISPR became a super fashionable topic," said Krzysztof Chylinski, who was a master's student at the time and developed an interest in CRISPR himself when he joined Charpentier's lab. He chalked up its popularity not just to the novelty of a complex adaptive immunity system in an organism no one had expected to contain one but also to the possible applications that occurred to many researchers.
The hypothesis of a bacterial immune system caught on, and a number of labs began to work on characterizing the CRISPR system and the CRISPR-associated, or Cas, proteins it requires.
Charpentier's microbiology lab in the Max Perutz Laboratories at the University of Vienna was one of them. The lab, which Charpentier had founded in 2002, worked on virulence factors in various pathogenic bacteria, including an RNA-regulated system in Streptococcus pyogenes. A bioinformatics screen they conducted in 2006 revealed that pyogenes too had a collection of clustered regularly interspaced repeats. They became interested in a noncoding RNA near the CRISPR locus that they thought might regulate the CRISPR system's activity.
Around the same time, Doudna, a structural biologist who had been focused on RNA interference in eukaryotic cells, learned about CRISPR through a conversation with Jillian Banfield, a colleague at the University of California, Berkeley. Intrigued by the similarities between eukaryotic RNAi and the CRISPR system, Doudna started investigating the structure and activity of Cas enzymes in Pseudomonas aeruginosa and, with collaborator Eva Nogales, published in 2011 a cryogenic electron microscopy structure of the E. coli CRISPR effector, a complex with many protein components.

Also in 2011, Chylinski was second author on a report from Charpentier's lab. They found that the noncoding RNA they had taken an interest in, dubbed the transactivating CRISPR or tracrRNA, had to be present for the S. pyogenes CRISPR RNA to mature into an active form and enable the pyogenes effector to cut the complementary target DNA.
There are some important differences between the E. coli and S. pyogenes CRISPR systems. Whereas E. coli and a majority of other bacteria have systems with many proteins guided by just one RNA, Charpentier's lab had realized that S. pyogenes requires two RNAs. Soon they would find that the system also involved only one effector protein.
The simplicity of the S. pyogenes system was tantalizing. Chylinski explained, "If in one system, you have like six proteins that are doing something, and in the other there is one, it must be one hell of a special protein."
The same type of scientist
Doudna and Charpentier met at a CRISPR conference in San Juan, Puerto Rico, in 2011 when the CRISPR field was still small enough to fit into a single room. They hit it off right away, both sharing a love of hard problems and careful investigation. Charpentier told a Nobel interviewer, "We are the same type of scientist, who want to see the details of the data."

in Doudna's lab and was co-first author
on the 2012 paper describing the mechanism
of Cas9.
The two labs came together for an often-described transcontinental and transoceanic collaboration. Though it involved just two labs, it took place in three locations: Berkeley, Vienna, and Umeå — a city in Sweden where Charpentier was in the process of moving her lab. Soon Chylinski and Martin Jinek, a postdoc in the Doudna lab, were hard at work using purified Cas9 protein and tracr and CRISPR RNAs to understand how Cas9 cuts target DNA.
The work went smoothly. None of the researchers recalled any extraordinary roadblocks.
"If it would be easy, everything would have been discovered already," Chylinski said. But in this project, the hurdles that arose were "just, you know, regular troubleshooting, and let's move forward."
The results, too, were clear. "Sometimes you work on systems, and it takes a long time to see what you would like to see — or the result is not black and white, it's light gray or dark gray," Charpentier explained to a Nobel interviewer. "Here, it was really white."
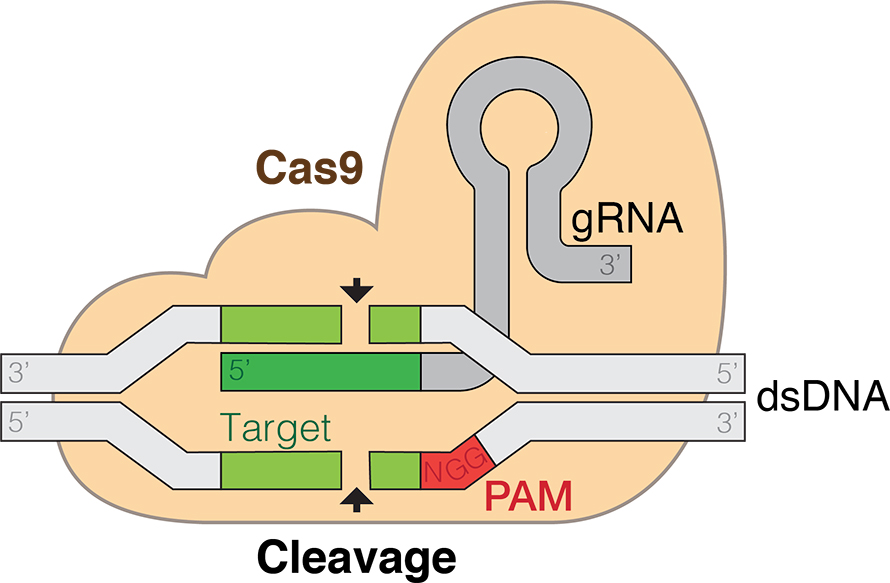
The team found that base pairing between the tracr and CRISPR RNA could lead Cas9 to cut a DNA target that matched the CRISPR RNA so long as the target DNA included a protospacer-adjacent motif nearby. They also demonstrated that a single engineered hairpin RNA could perform the same role — in other words, they established a two-component system for genetic engineering.

in Emmanuelle Charpentier's lab and was
co-first author on the 2012 paper describing
the mechanism of Cas9.
Because CRISPR had intrigued so many researchers, the team knew that they needed to publish soon to beat their competitors.
"Towards the end … it became kind of an around-the-clock operation," Jinek said. "We took advantage of the time difference." He would wake up to an email from Chylinski reporting on the day's results and would send a similar email before he headed home.
The work was published in Science in 2012. A year and a half later, they followed up, again in Science, with a detailed structural description of two Cas9 proteins from two bacterial species and how each of them coordinated DNA and RNA.
CRISPR's big break
The CRISPR field already was moving fast. Once the 2012 paper on reprogramming Cas9 was published, it exploded. Labs around the world picked it up rapidly. Already by the time of their second paper, Doudna, Charpentier and Nogales could cite seven major articles that had used CRISPR and Cas to induce double-stranded breaks in DNA, modulate transcriptional activity and modify eukaryotic genomes — and those were just the papers that already had been published.
"All of a sudden, tons and tons of labs could afford genome engineering as a very standard thing," Chylinski said. He fielded a slew of technical questions about the tool while finishing his Ph.D., and after graduation he took a job at a small nonprofit that provides technical services to help research labs implement the technology.

Doudna and Charpentier each launched biotechnology companies in 2013: Charpentier started CRISPR Therapeutics, while Doudna (who two years prior had started Caribou Biosciences based on her own lab's CRISPR work) teamed up with Feng Zhang of the Broad Institute and George Church from Harvard and MIT, along with several others, to start Editas Medicine. Later, overlapping applications for patents on Cas9-based genome editing pulled the UC system and the Broad Institute into a long-running, bitterly fought and well publicized dispute over ownership of the rights to use CRISPR–Cas9 genome editing in eukaryotic cells.

and a junior faculty member at Yale. "She
didn't need much mentoring, but everybody
can stand some advocacy," Steitz said.
Though it's still unclear who will win commercial rights to CRISPR–Cas9, in basic research the outcome is clear. "Everybody uses it," said RNA structural biologist Joan Steitz, who knew Doudna as a postdoc at the Unviersity of Colorado and a junior professor at Yale.
Societal implications
At this year's inaugural World CRISPR Day on Oct. 20 sponsored by the CRISPR service company Synthego, Fyodor Urnov shared a story about stepping out of his office at the IGI, which he co-leads with Doudna, for an espresso a week or two before the Nobel was announced.
He happened to be wearing a CRISPR T-shirt. The barista asked, "Do you do that genome editing thing?"
Yes, Urnov said. "In fact, I work with Jennifer Doudna."
Impressed, the barista gave him the coffee on the house.

Even before the prize, Doudna and Charpentier had reached international celebrity status, racking up a series of other high-profile recognitions. The pair shared a Breakthrough Prize in 2015, a Tang Prize in 2016 with CRISPR researcher Feng Zhang, a Japan Prize in 2017, a Kavli Prize in 2018 with Lithuanian CRISPR researcher Virginijus Šikšnys and a Wolf Prize earlier this year. In the wake of their discovery, both have founded institutes that they now lead: Doudna at the IGI and Charpentier, in 2018, at the Max Planck Unit for the Science of Pathogens in Berlin.
Of the two, Doudna seems more comfortable stepping onto the global scientific stage. Both have been involved in public conversations about the ethics of CRISPR, but Doudna has been more focused on bringing that conversation to nonscientists and has become the public face of the technology and its implications.
"For many scientists, this is a foreign area: we're not trained to talk about our work more broadly," she said. "Many of us prefer to do the next experiment rather than stepping out of the lab and thinking about how to communicate our science to non-specialists — but I do think it's essential to do that."
Urnov said during his World CRISPR Day talk, "I salute Jennifer, personally, for taking such a clear citizen-scientist role," adding that he hopes more scientists will take inspiration from her sense of civic responsibility.
A gifted explainer from the start, Doudna drew inspiration from the famous Asilomar conference in the 1970s, which established scientific guardrails around recombinant DNA. When planning a conference to discuss the implications of the CRISPR technology, which took place in Napa Valley in 2015, she invited two researchers who'd attended and organized the original, Paul Berg and David Baltimore.
More reading on how CRISPR was discovered:
"A Crack in Creation" by Jennifer Doudna with Samuel Sternberg
"The biology of CRISPR–Cas: Backward and forward" by Charpentier and colleagues
"A day with Jennifer Doudna" by Lisa Jarvis for Chemistry & Engineering News
"The quiet revolutionary" by Allison Abbott for Nature
"The promise and challenge of therapeutic genome editing" by Doudna
Many of those same researchers, including Doudna and Baltimore, were at the Second International Summit on Human Genome Editing in November 2018, when Chinese researcher He Jiankui announced the astonishing news that he had edited a receptor called CCR5 out of the genome of human embryos — and that edited twins recently had been born. The work was sloppily executed, and the scientific rationale for altering CCR5 was hazy; in January of this year He was sentenced to a three-year prison term. Scientists around the world renewed their calls for a moratorium on heritable editing, but He's announcement already had shown the limits of such appeals.
Many more CRISPR systems
While modification of human embryos sucks up a lot of the oxygen in public conversations about genome editing, Urnov may have spoken for more scientists than himself when he said at his World CRISPR day talk, holding a hand level with his eyebrows, "I've had it up to here" with discussions of germline editing.
The technology has been applied to many societal problems that are less fraught: agriculture, drug development and nonheritable gene therapy, to name a few. Most of the scientists interviewed for this story mentioned Victoria Gray, an American whose sickle cell anemia was cured by edits to stem cells extracted from and then transplanted back into her bone marrow.
CRISPR also has been used in basic research to understand topics as esoteric as the gene controlling snail shell spiral chirality. Meanwhile, the work to understand bacterial immunity goes on — and the applications for bacterial immunity effectors are far from limited to Cas9.
"Now we don't have just plain vanilla CRISPR/Cas9 that does DNA cutting," Jinek said. "We have base editors, we have prime editors, we have transcriptional control through CRISPR … all these applications that (are) still rooted in the basic molecular principle of having a guide RNA targeting double stranded DNA, but then apply all sorts of other molecular activities."
Five general types of CRISPR system are known, and researchers use them for different applications. Last year, for example, Doudna's lab published cryo-EM structures of CasX, which unlike Cas9 makes a staggered cut in the DNA, producing sticky ends. Other bacterial effector proteins cut RNA instead of DNA, produce cyclic second messenger molecules or simply bind specific sequences. With input from engineers, the list of applications for these protein functions continues to grow (see The many uses for CRISPR).

Researchers have scrambled to find and lay claim to other effector proteins of bacterial immunity by scanning and mining bacterial genomes. Jillian Banfield, the Berkeley microbiologist who introduced Doudna to CRISPR in the first place, runs a company called Metagenomi mining microbial genomes for novel gene-editing systems; the pharmaceutical company Bayer recently invested $65 million in the enterprise.
The future for CRISPR scientists
What could I possibly tell you about CRISPR technology that you haven't heard before? There have been about 48,000 research articles' worth of new findings this year alone. The technique has spurred close to a decade of explosive creativity in the fields of biotechnology, basic biological research and clinical medicine, and it shows no signs of slowing.
"It often feels like the genome engineering applications made possible by CRISPR are limited only by our collective imagination," Doudna wrote in her memoir, "A Crack in Creation." She used to keep a running list of organisms in which the technology had been used; that's no longer possible.
When asked what it was like to have been instrumental to such an important discovery, Chylinski answered with a puckish grin, "It's nice."
After a thoughtful pause, he added more seriously, "At the beginning it was almost intimidating how fast things were going. … In some places in the world, people are already getting experimental treatments partially based on something I did with my Ph.D. thesis."
Both Doudna and Charpentier have worked on a variety of topics over their careers, and as the CRISPR field expands, they've found new areas within it to work on.
Charpentier has had a peripatetic career, working in France, the U.S., Austria, Sweden and now Germany. "I could feel very quickly the right time to move on," she told an interviewer from The CRISPR Journal in 2019. She said she wanted "not to get stuck in a position where maybe I would not evolve the way I wanted to evolve."
Steitz observed that Doudna is "interested in all sorts of very specific experimental problems and questions within the whole big realm of RNA and all its functions in cells. She hasn't been monolithic."
Doudna agreed with that interpretation and said her work is all about a broader exploration of numerous interesting problems. "When I think about how I choose topics, it's more of a gut feeling," she said. "It's the things that I find interesting, or the things that I feel intrigued by, that tend to be the things I'm going to go after."
Yogurt research, broad applications
CRISPR stands for clustered regularly interspaced short palindromic repeats. Discovered in 1993 by Spanish scientist Francisco Mojica, these short repeating sequences interspersed with hundreds of diverse spacers appeared in the genomes of more and more different bacterial species as he continued to look. Researchers also found CRISPR-associated, or Cas, genes, that always seemed to travel with them.
In 2005, Mojica reported that many spacer sequences matched the genes of then-known phages, viruses that infect bacteria, which raised a provocative hypothesis: that the repeating elements could work as a bacterial form of adaptive immunity. The same year, researchers reported that bacteria with more CRISPR repeats tended to be resistant to more phages.
In 2007, researchers at the yogurt company Danisco, who were interested in reducing the company's loss of fermenting bacteria to phage infections, found that bacteria that acquired resistance to a phage isolate also acquired a spacer sequence matching a portion of that phage's genome.
This provided more evidence for the hypothesis that CRISPR works as a bacterial form of adaptive immunity. By absorbing snippets of phage genome and marking them between the molecular flags of repeated sequences, bacteria establish a sort of back file of invaders they've encountered. The RNA expressed from this genetic filing cabinet guides effector proteins in the Cas family to find and often destroy phage genomes that make it into the cell.
Evolution has generated immense diversity in CRISPR–Cas systems. "It's not like one mechanism describes everything," said Krzysztof Chylinski, who co-discovered the mechanism of the most famous CRISPR system as a graduate student. "There are two classes, six types and, I don't know, dozens of subtypes — and some uncharacterized systems."
CRISPR vs. COVID-19
In late September, researchers from Doudna's lab and collaborators at the Universities of California at Berkeley and San Francisco and the Chan Zuckerberg Biohub posted a preprint introducing a method for using a CRISPR-based system to diagnose COVID-19 without amplifying its nucleic acids. Instead of the famous Cas9 protein, which cuts DNA at a site of complementarity with a guide RNA, this assay uses the protein Cas13, another effector in the arsenal of bacterial CRISPR immune systems.
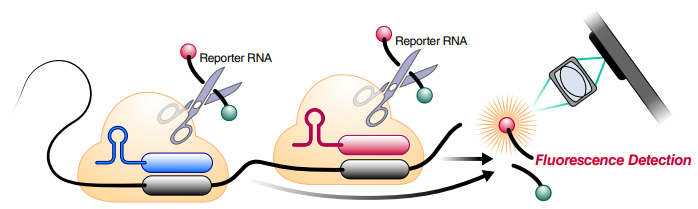
The technique is called specific high-sensitivity enzymatic reporter unlocking, or SHERLOCK. Cas13 responds to a guide–target match by cutting the target RNA — and, when purified, snowballing to cut every other RNA in the vicinity. That makes it great for detecting a small amount of nucleic acid. The assay has four parts: Cas13, its guide RNA, an RNA probe with a fluorescent moiety on one end and a quencher on the other end, and the patient sample. If base pairing happens between RNA in the patient sample and the guide RNA, then Cas13's wild reaction is triggered. Cutting the RNA probe separates the fluorophore from the quencher so that if SARS-CoV-2 RNA is present in the sample, fluorescence occurs.
In addition to adapting the chemistry from previous SHERLOCK applications, the team also came up with a detection box smaller than a shoebox with a stripped-down fluorescence excitation setup, which can be imaged using an ordinary cell phone running a custom app. Twenty-eight scientists including biochemists, virologists, biomedical engineers and physicists were involved.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in People
People highlights or most popular articles

de la Fuente honored for AI research
The award will support the development of an AI system called ApexMol, a 3D structure–informed, agentic large language model designed to create new biomolecules.

In memoriam: Peter Roepstorff
He was a leading researcher in biological mass spectrometry, mapped protein function in living organisms and was an ASBMB member for 19 years.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

ASBMB members receive RNA Society awards
The RNA Society awards Brenda Bass, Can Cenik and Karin Musier–Forsyth for their achievements in RNA research and innovation. Winners will be recognized at the closing awards ceremony of the RNA 2026 annual meeting.

In memoriam: Richard L. Cross
He studied the enzymatic mechanisms of ATP synthase and served on the editorial board of the Journal of Biological Chemistry for 24 years.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.


