Merging biochemical and analytical training
Recent advances in molecular and biochemical methods such as mass spectrometry and high-throughput sequencing have accelerated the rate of scientific discovery and exponentially increased the volume of data that a single study can generate. The COVID-19 pandemic has stimulated researchers to integrate genome sequencing, automatized large-scale testing, rapid and efficient sharing of information, modeling, and computational studies to generate evidence-based solutions to global problems.
We also have been challenged to be more creative and flexible in our approach to the way we work and teach. For some scientists, this has meant increasing our reliance on computer networks and computational methods to compensate for limited access to the laboratory bench. In other words, the pandemic revealed that while it is critical for us to specialize and have depth of knowledge in some domains, it is also essential that we cultivate some breadth in our skill set.
A need for programming literacy
Question-driven research often requires the analysis of large volumes of heterogeneous data to generate accurate and comprehensive answers. Often, these data must be integrated into predictive models. For example, in my laboratory we integrate headspace volatiles analysis, transcriptomics and electrophysiological recordings to understand the chemical basis of mosquito–host interactions.
Integrated multidisciplinary research is not new and not limited to the fields of biochemistry and molecular biology. In both academia and industry, scientists at all levels need to be able to work at the bench and operate advanced scientific equipment as well as process, wrangle, integrate and analyze heterogeneous data. Our colleagues in neuroscience combine behavioral data with neural activity recordings, functional imaging studies, gene expression profiles, computational approaches and mathematical modeling. Ecologists integrate temporal population census data with climatic and geographic information to discern how organisms interact with their environment. As these multidisciplinary approaches spread, we need to train what data scientists refer to as “pi-shaped” researchers: investigators possessing a broad understanding of the sciences supported by a deep knowledge of their specific area of expertise and a foundation in data science. The presence of these π-shaped experts on problem-solving teams facilitates the process of data collection, visualization and interpretation by freeing these teams from communication gaps between topic expertise and data scientists.
Training π-shaped researchers

Data science requires both computer literacy and familiarity with a programming language. While some students are curious about programming and learn how to code in Python or R on their own, others don’t have this exposure to data wrangling unless they are part of the roughly 54% of undergraduates (averaged across science, technology and engineering and mathematics disciplines) who participate in extracurricular research. At most colleges and universities, an extensive set of prerequisites is required for students to take advanced specialty classes in computer science and statistics — a challenge for biochemistry students who already have full schedules. In most bench and field science majors, computational training is optional and not part of a student’s core training. As a result, students have difficulty identifying how knowledge acquired in a statistics or computer science course can be applied to, for example, biochemistry.
In the biochemistry program at Virginia Tech, we developed a new course that exposes biochemistry majors to coding as they analyze large data sets relevant to the concepts and topics developed in class — for example, chemical communication. Students analyze large data sets collected in the instructor’s laboratory, ranging from electrophysiological recordings of olfactory neurons to gas chromatography–mass spectrometry analyses of the chemical composition of plant and human scent samples. In addition to programming in the open source language R, they work in teams to clean, wrangle, visualize and interpret data. They manipulate inferential statistics and use multivariate analysis and machine learning while answering biochemical and biological questions.
Copy, paste and tweak
If you were to type, “La mer, la vaste mer, console nos labeurs!” (from Charles Baudelaire’s poem “Moesta et Errabunda”) on your computer, would you have written one line of French poetry? Yes. Does this mean that you now are a poet or know how to communicate in French? Not exactly, right? The same applies to learning a programming language. Providing students with functional scripts does guarantee that they will produce an anticipated output. However, such assistance reduces the likelihood that they will be able to then tackle a slightly different problem. On the other hand, expecting non–data science students to become programming experts in a single semester is unrealistic.
Through a compromise approach, our students can acquire a working understanding of the programming relevant to their area of study. By working with data that students can relate to and that is directly relevant to the topic of the course, we offer them an opportunity to leverage lecture content and reading materials to identify the biochemical problem they are trying to solve. The central pedagogical objective is to foster students’ familiarity with key coding concepts and terms and to develop their ability to identify code syntax and structures that they can adapt to fit their needs and solve the biochemical problem.
Optimizing online
How does this look in the classroom? Before the pandemic, students worked side-by-side in small groups to brainstorm, code and debug while the instructor and teaching assistant moved between groups to provide individualized teaching. Physical distancing requirements have disrupted these activities, but online solutions exist that emulate these interactions.
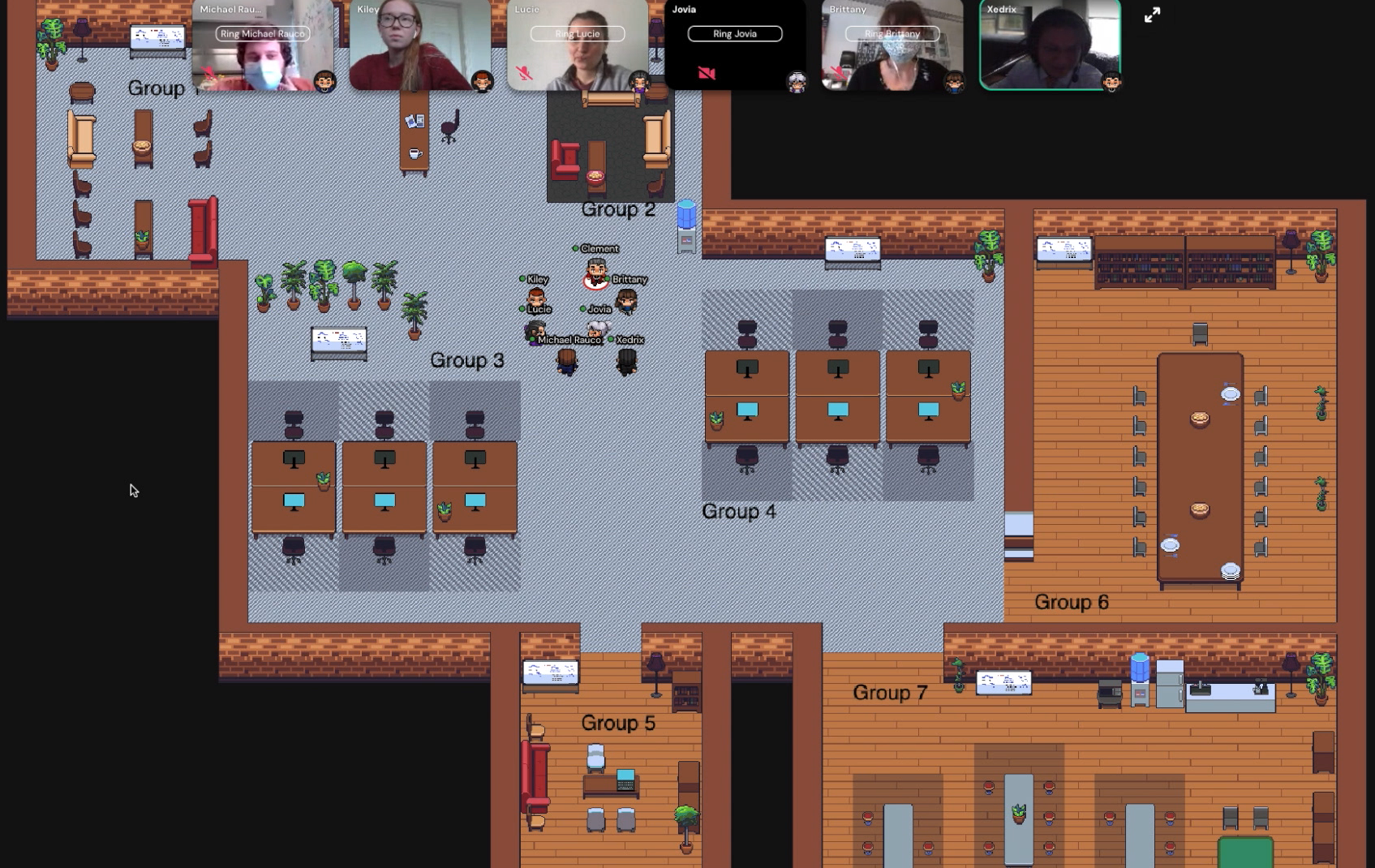
During the spring semester of 2021, our class met in a virtual classroom on Gather.com, a video call platform. It is similar to Zoom or Teams, but each participant has an avatar that can move around the virtual classroom. Students worked collaboratively on their codes using platforms such as Google Drive and GitHub. With these tools, they were able to share their work with the instructors and get feedback and personalized help, and instructors were able more readily to comment, edit students’ code in real time and explain core coding concepts. We recorded lectures and group activities so students with added responsibilities (such as parenting), disabilities or scheduling conflicts can come back to the material later.
Career benefits
Looking forward to the fall semester, now is a good time to reimagine a post-pandemic version of this new course in which analytical training can be even better integrated with core biochemistry education. Returning to in-person teaching should increase student engagement and mitigate some of the inequalities arising from their work-from-home environments. They still will be able to share and exchange data and work collaboratively online.
We have a unique opportunity to prepare undergraduates for professional scientific collaborations that are often long-distance, if not international. As we observed during the pandemic, online resources for collaborative work offer a remarkable medium to provide individualized feedback to students, bringing coursework one step closer to the one-on-one training students would get in a laboratory. By recording and sharing lectures and discussions via online platforms, we are able to reach students with varied learning styles and needs.
A foundation in data science tailored for biochemists and life scientists will give students an edge when applying to graduate or medical school or entering the job market. By exposing students to the use of online resources for collaborative work, we help them to hit the ground running when they move on to the next step of their training or the first step of their professional life.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Opinions
Opinions highlights or most popular articles

Learning can be fun: Gaming anatomy and physiology
Instructors explore how gamification and active learning transform student engagement and retention. They convey how emotion, interaction and design can make even rigorous subjects more effective and memorable.

Mentorship and uncertainty: Lessons from Telemachus
A biochemistry educator reflects on mentorship through the Greek story of Telemachus, showing how embracing uncertainty, failure and curiosity can transform teaching.

Embracing the twists and turns along the educator pathway
A biochemistry educator reflects on the challenges of early faculty life, describing how evidence-based teaching, cross-disciplinary collaboration and classroom challenges shaped her growth.

Redesigning with students in mind
Assistant professor reflects on how the shift to online teaching revealed gaps in points-based grading and led to a redesign centered on transparency and student growth.

Teaching beyond information transfer
Educator reflects on moving beyond lectures to create a biochemistry classroom centered on engagement, transparency and student ownership, showing how small shifts like “student hours” and active learning can transform understanding.

Mayday! Lessons from cellular dysfunction and group work dynamics
An upper-level biology course revealed that strong science doesn’t guarantee strong teamwork. One instructor shares how failed group dynamics reshaped their approach, leading to more structured, collaborative and effective student learning.

