Winners of the ‘aha moments’ essay contest
To celebrate our three journals going open access, we invited readers to share their moments of discovery in science. Here are the first, second and third place winners.
FIRST PLACE
Finding a common ancestor
By Richard Ludueña
As a child, I loved science fiction. I always have liked history, and I especially was drawn to stories about time travel, because I loved to imagine looking far back into the past.
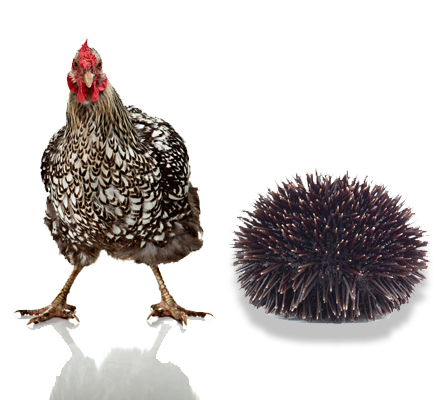

My aha moment came in 1972 when I was a graduate student in Dow Woodward's laboratory at Stanford determining the first 25 amino acids in the sequences of alpha- and beta-tubulin from chickens and sea urchins. At that time, very little was known about tubulin sequences.
The results came off the sequencer around Christmas Day. I realized that chicken and sea urchin α-tubulin were about 95% identical and that the same was true of the β-tubulins.
This meant that I had just learned something very intimate about the common ancestor of vertebrates and echinoderms, an organism whose appearance and morphology were unknown and only could be guessed at. I felt like I was looking perhaps 700 million years into the past. I had no idea what that animal looked like, but I knew something about its tubulin.
I also observed that α- and β-tubulin were themselves about 40% to 45% identical, so I now perhaps was seeing more than 2 billion years into the past to the time of the first eukaryotes, one-celled organisms that lived on a planet that would be unrecognizable to us.
This was one of the most exciting moments of my career as a biochemist and the closest I ever came to realizing my childish fantasy of time travel. I think a child lives inside every scientist, a child whose curiosity is challenged by mysteries, who wants to make visible the invisible and to magnify the microscopic and who wants to look back into time as far as the origins of everything — of cells, life, the earth and the universe.
SECOND PLACE
Dreaming of Western blots
By Mindy Engevik
It was a cold night; the wind was blowing, and the branches of a nearby tree were scratching against the roof of my apartment. I was in bed huddled under a big down comforter, which enveloped my whole body like an amoeba. I was deep in REM sleep, dreaming of Western blots. Horrible Western blots … the bane of my existence.
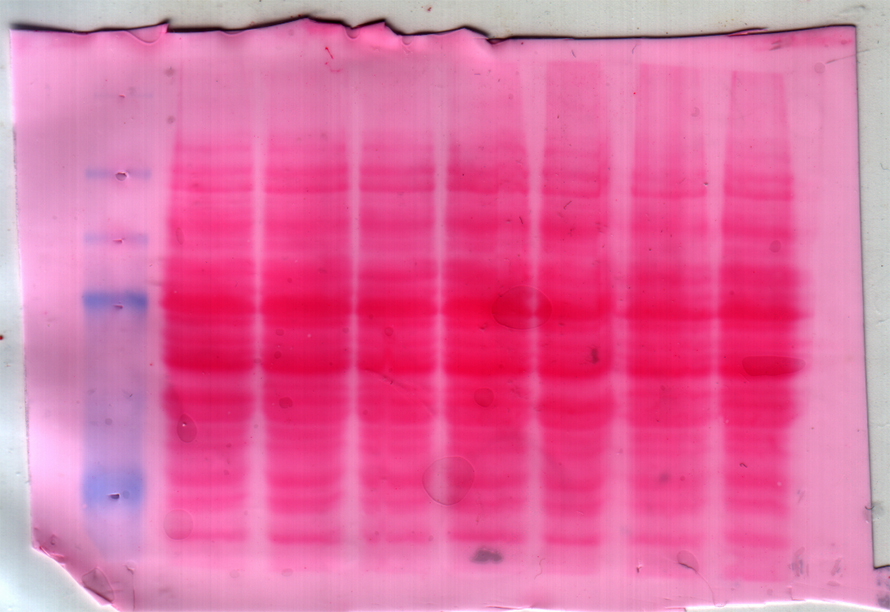

As a graduate student, I was fascinated by the beautifully orchestrated events of ion transport of the intestine. I gloried over ion transporter mRNA, enjoyed functional assays in tissue cultures, reveled in measuring fecal ion concentrations by flame photometry — but then I encountered the dreaded Western blot.
I needed to measure the protein levels of the sodium–hydrogen exchanger isoform 3 and the colonic HK-ATPase in mouse colon by Western blot. Day after day, I had been troubleshooting these Western blots. I had tried optimizing the denaturing steps, removed the denaturing step, changed the blocking buffer, modified the run times … I had tried everything I could think of and everything my helpful colleagues had recommended. But to no avail — the Western blot had won.
But that night, in my dreams, I went over the Excel sheet provided by my PI that provided a handy plug-in for calculating the amount of protein to add to the Western blots. I dutifully added my dream concentrations and realized … the math was wrong.
I bolted awake, ran to my laptop, checked the Excel sheet and voila — the Excel sheet math equation was incorrect!
That morning, I went to lab, added the correct amount of protein, and this time, the Western blot worked. I felt like a conquering hero. I guess dreams really do come true.
THIRD PLACE
Beauty in brown
By Kazuhiko Igarashi
The laboratory where I started as a new faculty member, led by Norio Hayashi and Masayuki Yamamoto, focused on erythroid cells: Colleagues were working on the regulation of heme synthesis during erythropoiesis, and I was trying to identify transcription factors that would regulate globin genes by binding to their superenhancers, the locus control region, together with two graduate students.
One student, Ken Itoh, got two promising candidate cDNA clones in two hybrid screenings using MafK, one of the subunits of nuclear factor erythroid 2, as a bait. The other student, Tatsuya Oyake, expressed one clone in E. coli for purification. When he came out of the cold room, he was disappointed, thinking that his purification was a total failure. The protein solutions looked brown.
We were troubleshooting in front of the cold room, with the tubes in his ice box. Then our lab head, Norio Hayashi, who has a long track record in enzymes of heme synthesis, happened to walk by. It did not take a second before he saw the tubes and suggested carefully, "This may be a heme protein."
A beauty emerged: Heme, which is massively synthesized during erythropoiesis, would regulate the activity of this transcription factor, BACH1, coordinating heme synthesis and globin gene expression.
As so many scientists know, the simplicity of a model does not guarantee a simple way forward. For us, it took another two decades to come to a reasonably correct answer on the function of the heme–BACH1 axis in erythropoiesis. Various functions of BACH1 and the other clone, BACH2, which also is regulated by heme, have emerged in not only erythropoiesis but also iron and heme metabolism, immune response and cancer progression.
If Tatsuya had discarded the tubes out of disappointment or Professor Hayashi had not seen those tubes, what would we be doing today?
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Opinions
Opinions highlights or most popular articles

Learning can be fun: Gaming anatomy and physiology
Instructors explore how gamification and active learning transform student engagement and retention. They convey how emotion, interaction and design can make even rigorous subjects more effective and memorable.

Mentorship and uncertainty: Lessons from Telemachus
A biochemistry educator reflects on mentorship through the Greek story of Telemachus, showing how embracing uncertainty, failure and curiosity can transform teaching.

Embracing the twists and turns along the educator pathway
A biochemistry educator reflects on the challenges of early faculty life, describing how evidence-based teaching, cross-disciplinary collaboration and classroom challenges shaped her growth.

Redesigning with students in mind
Assistant professor reflects on how the shift to online teaching revealed gaps in points-based grading and led to a redesign centered on transparency and student growth.

Teaching beyond information transfer
Educator reflects on moving beyond lectures to create a biochemistry classroom centered on engagement, transparency and student ownership, showing how small shifts like “student hours” and active learning can transform understanding.

Mayday! Lessons from cellular dysfunction and group work dynamics
An upper-level biology course revealed that strong science doesn’t guarantee strong teamwork. One instructor shares how failed group dynamics reshaped their approach, leading to more structured, collaborative and effective student learning.



