At the interface
Sphingolipids, or SLs, have emerged as critical players in membrane stability and as essential signaling molecules. SLs range from abundant species involved in maintaining membrane integrity, such as sphingomyelin, to scarce and potent signaling species, such as sphingosine-1-phosphate, or S1P. S1P mediates critical signaling functions through interaction with its cognate G-protein coupled receptors in development and in several disease states. In recent decades, many resources have been devoted to understanding how S1P generation is regulated.
S1P can be generated through the action of two sphingosine kinase isoforms. The more commonly expressed of the two is sphingosine kinase 1, or SK1. This enzyme has garnered attention as a potential therapeutic target, as it often is upregulated in diseases such as cancer. To gain access to its substrate, sphingosine, and to release its product, S1P, SK1 must interact directly with membranes. However, how SK1 achieves membrane binding has been contested in the literature. How does a cytosolic lipid-metabolizing enzyme without any lipid-binding domains interact with membranes to access its substrate?
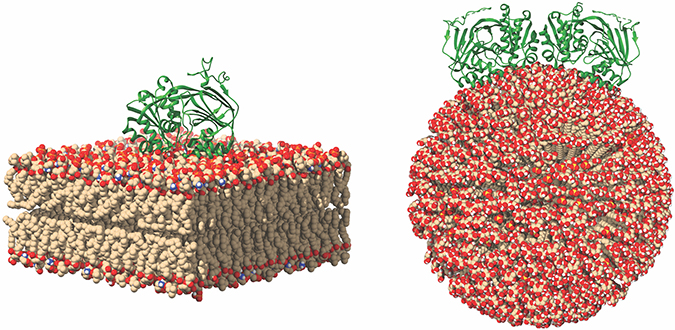 This image of SK1 binding to membranes shows potential SK1 membrane binding poses for SK1 in the presence of flat and curved membranes. Michael Pulkoski-Gross
This image of SK1 binding to membranes shows potential SK1 membrane binding poses for SK1 in the presence of flat and curved membranes. Michael Pulkoski-Gross
Previously, researchers thought SK1 translocation was dependent on other proteins. However, recent data show how SK1 can bind directly to membranes. SK1 possesses an intrinsic interface composed of two motifs: one electrostatic motif and one hydrophobic motif. Using biochemical methods, we found that these two motifs are necessary for membrane interaction, thus implicating their function as a single entity. Using hydrogen deuterium exchange mass spectrometry, we confirmed that SK1 employs a single contiguous interface that contains the two motifs. In cancer cells, disruption of this interface causes loss of membrane association and decreases SK1 activity. Past research has shown that interaction with membranes is critical for mediating SK1-dependent biologies including tumor cell invasion and endocytosis. This could provide a new avenue for targeting SK1 in diseases. Inhibition of membrane binding would deny SK1 access to its substrate, thereby inhibiting all activity.
Recent research shows an important role for SK1 in endocytic trafficking. SK1 presence at endocytic membranes would require membrane binding and curvature sensing. However, how SK1 can do this is largely unknown. Analysis of the atomic structure of SK1 revealed a potential dimerization interface. Such dimerization would align the membrane-binding interface of each SK1 monomer. This would strengthen the interaction and potentially allow for physical curvature sensing by SK1. This remains to be validated, but it’s an exciting hypothesis. Another way to potentially inhibit activity would be to inhibit dimerization, if that is required for activity and membrane binding.
Significant strides have been made in understanding the structure and function of many SL metabolizing enzymes, and exciting questions remain to be answered, especially for SK1. How can SK1 decipher the difference between the charges of different anionic phospholipids? How does the catalytic cycle progress once SK1 is at the membrane? Is dimerization required for membrane binding/curvature sensing? What role is the hydrophobic patch playing in curvature sensing?
Biophysical, biochemical and structural research will reveal the secrets of how SL enzymes work and how they might be exploited for therapeutic development.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.

How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.
The molecular orchestra of memory
Calcium, calmodulin and calcium/calmodulin-dependent kinase II form a molecular axis that turns fleeting neural activity into lasting memories. New research shows how memories are stabilized, and possibly even protected or repaired.


