From the journals: June/July 2018
We offer a selection of recent papers on a variety of topics from the Journal of Biological Chemistry, the Journal of Lipid Research and Molecular & Cellular Proteomics.
Looking at cells, living and dead
 Amir Ata Saei and colleagues at the Karolinska Institute assessed the proteome of adherent and dying cultured cancer cells after treatment. In this scanning electron micrograph, a HeLa cell in the midst of apoptosis begins to detach from its substrate. Courtesy of the National Institutes of Health Researchers at the Karolinska Institute were using a proteomic screen to determine the mechanism by which cancer drug candidates killed cells when they realized a potential flaw in the standard experimental design. Amir Ata Saei and colleagues usually treated adherent cancer cells with a compound and then rinsed away any cells that detached and measured how the proteome had changed in cells that survived. But, the researchers reasoned, the dying cells were responding the most strongly to the experimental drug. Shouldn’t their responses be examined?
Amir Ata Saei and colleagues at the Karolinska Institute assessed the proteome of adherent and dying cultured cancer cells after treatment. In this scanning electron micrograph, a HeLa cell in the midst of apoptosis begins to detach from its substrate. Courtesy of the National Institutes of Health Researchers at the Karolinska Institute were using a proteomic screen to determine the mechanism by which cancer drug candidates killed cells when they realized a potential flaw in the standard experimental design. Amir Ata Saei and colleagues usually treated adherent cancer cells with a compound and then rinsed away any cells that detached and measured how the proteome had changed in cells that survived. But, the researchers reasoned, the dying cells were responding the most strongly to the experimental drug. Shouldn’t their responses be examined?
In a study published in Molecular & Cellular Proteomics, the researchers compared proteomes of cultured cancer cells that had survived with those that had not after treatment with various drug candidates. Using drugs with known targets, they found that including dying cells improved the accuracy of target identification. To their surprise, the researchers also found that some proteins were upregulated in all detached and dying cells, regardless of the drug they used. They propose that these proteins, which previously had not been linked to cell death, may be cellular decision makers
and promising chemotherapeutic targets.
Diet regulates a metabolite
(but not in the brain)
Ketogenic diets, which reduce carbohydrate intake and prompt the body to rely on fat-derived ketone bodies instead, are a popular treatment for epilepsy and thought to have neuroprotective effects on some other diseases. Mild caloric restriction also is believed to protect neurons. Researchers aren’t sure of the exact molecular mechanism of these diets, but Svenja Heischmann and colleagues at the University of Colorado in Denver have taken a step toward characterizing their effects on the brain.
In a study reported in the Journal of Lipid Research, researchers conducted a metabolomics analysis of both the plasma and brain tissue of mice eating normal or ketogenic chow. They subdivided each diet group into mice eating their fill or eating a restricted amount of chow. The researchers found that, in the bloodstream, kynurenine metabolism changed dramatically. Kynurenine, made from the amino acid tryptophan, can be converted into vitamin B3 or several other metabolites with effects on neurons. However, in the brain, the level of kynurenine changed relatively little.
The research suggests that, while tryptophan degradation is a target of the ketogenic diet, changes in plasma metabolism may not always cross the blood-brain barrier. The researchers intend to explore other metabolic changes in future publications.
Caspase switch shows how to kill a killer
 This ribbon diagram represents a structure of caspase-3 (Protein Data Bank number 1qx3).courtesy of Jawahar Swaminathan/European Bioinformatics Institute The mammalian protein caspase-3 is an executioner with several faces. The protease initiates various processes involved in breaking down cells during apoptosis; for this reason, its regulation is important for controlling cancers. But low levels of caspase-3 activity are required for normal development.
This ribbon diagram represents a structure of caspase-3 (Protein Data Bank number 1qx3).courtesy of Jawahar Swaminathan/European Bioinformatics Institute The mammalian protein caspase-3 is an executioner with several faces. The protease initiates various processes involved in breaking down cells during apoptosis; for this reason, its regulation is important for controlling cancers. But low levels of caspase-3 activity are required for normal development.
Researchers from North Carolina State University and the University of Texas at Arlington have gained new insights into how caspase-3 is regulated, potentially providing opportunities for fine-tuning its activity. Postdoctoral research associate Melvin Thomas III led the study, which was published in the Journal of Biological Chemistry.
“Caspase-3 has been well-characterized in apoptosis; (it starts) the process of programmed cell death,” Thomas said. “But over the past decade, there’s been a lot of evidence that shows that caspases in general, but caspase-3 specifically, is involved in other non-apoptotic functions, such as erythroid differentiation, macrophage differentiation, neuron pruning and a variety of other developmental phenotypes.”
Thomas, working in Clay Clark’s laboratory at North Carolina State, investigated how caspase-3 was regulated allosterically by phosphorylation. He saw that phosphorylation of different sites in the protein had different effects on its activity. In particular, phosphorylation of a site conserved across all apoptotic caspases reduced caspase-3 activity, whereas phosphorylation of a site specific to mammalian caspase-3 completely abolished its activity. Thomas hypothesizes that these two different types of switches allow the enzyme to be regulated to the right level of activity for different functions.
“You can use these posttranslational modifications to either decrease activity of the enzyme or, in the case of the kill switch, decrease it to zero,” Thomas said. “(The cell can) bring the total level of caspase activity below a threshold to allow for these developmental phenotypes without committing the cell to death.”
Sugars can slow diabetes drugs
Liraglutide is an analogue of glucagonlike peptide-1, or GLP-1, that is used in the treatment of Type 2 diabetes and cardiovascular diseases. Liraglutide binds to human serum albumin, which helps it traffic to the GLP-1 receptor. Angélique Gajahi Soudahome and colleagues at Université de La Réunion showed that liraglutide binding to albumin decreased with increasing albumin glycation. As albumin glycation increases with diabetes progression, this factor might affect the therapeutic efficacy of the drug. The study was published in the Journal of Biological Chemistry.
Engineering a better matriptase inhibitor
Matriptases are membrane-anchored serine proteases. When dysregulated, matriptase activates growth factors involved in cancer proliferation and metastasis. In healthy tissue, matriptase is inhibited by hepatocyte growth factor activator inhibitor type-1, or HAI-1. Aaron C. Mitchell and colleagues at Stanford University engineered a matriptase inhibitor based on HAI-1 in which an inactive domain was replaced with a second matriptase-binding domain, resulting in greater binding activity and inhibition of growth factor activation in lung, breast and prostate cancer cells. The study was published in the Journal of Biological Chemistry.
The structure of bacterial sensitivity to copper
 Copper is a ubiquitous antibiotic. Some door handles in hospitals are made of copper alloys to impede bacterial growth, and within the human body, the innate immune system mobilizes copper to sites of infection to help kill bacteria. For some pathogenic bacteria, therefore, the ability to tolerate copper is key to their ability to infect humans.
Copper is a ubiquitous antibiotic. Some door handles in hospitals are made of copper alloys to impede bacterial growth, and within the human body, the innate immune system mobilizes copper to sites of infection to help kill bacteria. For some pathogenic bacteria, therefore, the ability to tolerate copper is key to their ability to infect humans.
Researchers at the University of Queensland and Griffith University in Australia investigated the structural and redox biology of a bacterial protein involved in copper tolerance. Their findings, published in the Journal of Biological Chemistry, could spur the development of new tools to combat antibiotic-resistant infections.
Jennifer Martin of Griffith University, who oversaw the research, studies the suppressor of copper sensitivity, or Scs, family of proteins, which are critical for copper resistance in pathogens such as Proteus mirabilis, which causes urinary tract infections and hospital-acquired infections. Scs proteins are diverse; although they are related to the well-characterized disulfide bond-forming proteins found in Escherichia coli, the mechanisms by which they allow bacteria to tolerate copper are poorly understood.
The new study, led by University of Queensland graduate student Emily Furlong, focused on ScsB-alpha and ScsC in Proteus mirabilis. The biochemical data showed that the two proteins formed a redox relay in the bacterial cells. The structural data showed that ScsB-alpha interacts with ScsC via two immunoglobulinlike folds, a structure also found in an E. coli disulfide bond-forming protein.
“What was quite surprising was the structure of ScsB-alpha,” Furlong said. “Even though there’s a very low sequence identity (between ScsB-alpha and the E. coli protein), only 12 percent, the structural similarity is quite high.”
Understanding these structure-function relationships in Scs proteins will allow them to be targeted therapeutically. Disarming bacteria by making them newly sensitive to copper represents a new approach to treating infections. But questions remain as to how this approach might work in clinical settings.
“We’re still at the stage of trying to find small molecules to inhibit the (disulfide bond-forming) proteins, and we haven’t got to that stage with the Scs proteins, because it’s still fairly early in the piece,” Martin said. “There’s a lot of unknowns. What happens if you have an established infection and then you take a drug that disarms bacteria? Will that have an impact on the course of infection? Will it reduce the propensity of bacteria to develop resistance? Can it be used in combination with antibiotics that are currently unusable because of resistance? These are all questions that we don’t yet know the answers to.”
Monitoring proteomics in real time
Nothing is more frustrating than being forced to throw out data. After taking the time to prepare samples, take measurements and analyze the data, a researcher may realize that a problem with the samples or an instrument compromised the results. With luck, the realization happens early — but work still must be repeated. In long-running clinical proteomics studies, sometimes the realization doesn’t occur until all of the data are pooled, and the cost can be high.
In a recent article in Molecular & Cellular Proteomics, researchers at the Pacific Northwest National Laboratory working on a longitudinal study of the development of Type 1 diabetes announced that they have developed a tool that can tell in real time whether the quality of proteomics data has dropped. Bryan Stanfill and colleagues developed an algorithm that considers statistical features of mass spectrometry data to make a constantly adjusted model for comparison to a baseline of known high-quality data. By intentionally manipulating the instrument — for example, by changing an ion lens setting — they showed that the algorithm could identify times when the instrument needed to be recalibrated. The software also helped to flag points when unscheduled cleaning and maintenance were required. The software, called QC-ART, is freely available on the software sharing site GitHub.
HDAC’s role in endosomal pH
The endolysosomal system is characterized by a precisely controlled pH gradient along the pathway, ending with the acidic lysosome. Defective pH regulation in this pathway is associated with numerous disorders including neurodegenerative diseases. Hari Prasad and Rajini Rao from Johns Hopkins University performed a meta-analysis of factors affecting the regulation of an endosomal ion exchanger. They found, conserved across yeast, flies and mammals, that these exchangers (and therefore endosomal pH) were regulated transcriptionally by inhibition of a histone deacetylase in response to nutrient limitation. Pharmacologically increasing expression of the histone deacetylase corrected endosomal pH and improved clearance of amyloid proteins in a cell model of Alzheimer’s disease. The research was published in the Journal of Biological Chemistry.
Brown tissue whitening causes cell death, inflammation
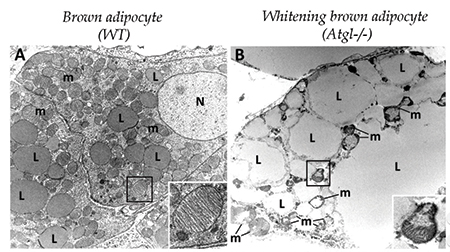 A brown adipocyte (left) has numerous small lipid droplets and healthy mitochondria (inset). In a whitening brown adipocyte (right), lipid droplets are much larger, and mitochondrial structure is disrupted.courtesy of KOTZBECK ET AL/UNIVERSITY OF GRAZ There’s more than one type of fat cell. Besides the white fat that stores triglycerides in lipid droplets in preparation for lean times later, mammals also have heat-generating brown fat, which acts more like a radiator than a storage closet. Brown fat cells are smaller, with more abundant mitochondria than white fat cells, and they hold a lot fewer lipids. In many models of obesity, brown adipose tissue converts to white tissue, with changes in the morphology and function of the cells.
A brown adipocyte (left) has numerous small lipid droplets and healthy mitochondria (inset). In a whitening brown adipocyte (right), lipid droplets are much larger, and mitochondrial structure is disrupted.courtesy of KOTZBECK ET AL/UNIVERSITY OF GRAZ There’s more than one type of fat cell. Besides the white fat that stores triglycerides in lipid droplets in preparation for lean times later, mammals also have heat-generating brown fat, which acts more like a radiator than a storage closet. Brown fat cells are smaller, with more abundant mitochondria than white fat cells, and they hold a lot fewer lipids. In many models of obesity, brown adipose tissue converts to white tissue, with changes in the morphology and function of the cells.
In a recent paper in the Journal of Lipid Research, Petra Kotzbeck, Antonio Giordano and colleagues investigated what happens to brown fat cells after whitening. The researchers, based at the University of Graz, Austria, and the University of Ancona, Italy, found that whitened brown adipocytes enlarged by addition of lipids were more likely to die than white adipocytes of a comparable size. Whitened adipose tissue also had more macrophages, presumably there to clean up the dead cells, and more inflammation under way. The vulnerability of whitened brown adipocytes may explain why gaining fat in the abdomen, where most whitened brown fat is located, is worse for your health than gaining subcutaneous fat.
An oncogene ally against multiple myeloma
Transmembrane prostate androgen induced protein, or TMEPAI, is a poorly understood oncoprotein that is overexpressed in solid cancers but not in liquid cancers such as leukemia and lymphoma. Yanyun Du and colleagues at Suzhou Municipal Hospital and Soochow University in China showed that TMEPAI actually may protect against multiple myeloma, a plasma cell cancer. The authors found that TMEPAI overexpression induces multiple myeloma cell apoptosis through TMEPAI’s effect on a myeloma-promoting transcription factor. The study was published in the Journal of Biological Chemistry.
Diabetes lessons from diverse mice
Type 2 diabetes risk is influenced significantly by genetics, but most previous diabetes research in mice has focused on only a few mouse genotypes, limiting mechanistic insight into the genetic determinants of diabetes. In a study published in the Journal of Biological Chemistry, Kelly Mitok and colleagues at the University of Wisconsin–Madison examined diabetes-related metabolic traits and pancreatic islet proteomics in the genetically diverse Collaborative Cross panel of mice. They uncovered mouse strain-specific differences in diabetic phenotypes and pancreatic islet proteomic profiles. They also showed that in only one strain, pancreatic synthesis of dopamine inhibited insulin secretion, demonstrating the importance of a genetically diverse reference panel.
Syntaxin 17 promotes lipid droplet formation
Cells store energy in lipid droplets, and many such droplets are made in the liver, which plays an important role in coordinating fat metabolism. As new lipid droplets form within the endoplasmic reticulum, acyl coA synthetase 3, or ACSL3, is indispensable for helping them mature. ACSL3 turns free fatty acids into the neutral lipids that fill the lipid droplet.
In a recent article in the Journal of Lipid Research, Hana Kimura and colleagues studying droplet synthesis at Tokyo University of Pharmacy and Life Sciences in Japan report that the binding and scaffolding protein Stx17 is required to move ACSL3 to the nascent lipid droplet at mitochondria-associated membranes within the ER. This new role may explain why Stx17 is expressed abundantly in the liver and adipocytes.
How soy resists salt
How will crops cope with rising sea levels and increasing salt in the water table as the climate changes? In a recent study in Molecular & Cellular Proteomics, researchers at Hangzhou Normal University studied soybeans’ response to salt stress. Salt disrupts mitochondria and increases the reactive oxygen species in the plant cells. Therefore, increasing production of antioxidant molecules like flavonoids may protect the plant. By combining phosphoproteomics and metabolomics, Erxu Pi and colleagues described a salt-stress signaling pathway in soybean roots that increases flavonoid synthesis and improves salt tolerance.
 Sasha Mushegian is scientific communicator for JBC.
Sasha Mushegian is scientific communicator for JBC. Laurel Oldach is a communications intern at the ASBMB.
Laurel Oldach is a communications intern at the ASBMB.Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Hope for a cure hangs on research
Amid drastic proposed cuts to biomedical research, rare disease families like Hailey Adkisson’s fight for survival and hope. Without funding, science can’t “catch up” to help the patients who need it most.
Before we’ve lost what we can’t rebuild: Hope for prion disease
Sonia Vallabh and Eric Minikel, a husband-and-wife team racing to cure prion disease, helped develop ION717, an antisense oligonucleotide treatment now in clinical trials. Their mission is personal — and just getting started.

Defeating deletions and duplications
Promising therapeutics for chromosome 15 rare neurodevelopmental disorders, including Angelman syndrome, Dup15q syndrome and Prader–Willi syndrome.
Using 'nature’s mistakes' as a window into Lafora disease
After years of heartbreak, Lafora disease families are fueling glycogen storage research breakthroughs, helping develop therapies that may treat not only Lafora but other related neurological disorders.

Cracking cancer’s code through functional connections
A machine learning–derived protein cofunction network is transforming how scientists understand and uncover relationships between proteins in cancer.

Gaze into the proteomics crystal ball
The 15th International Symposium on Proteomics in the Life Sciences symposium will be held August 17–21 in Cambridge, Massachusetts.

.jpg?lang=en-US&width=300&height=300&ext=.jpg)
