JBC: Series explores interactions between microbes and the gut
The human gut is teeming with microbes that are beneficial and sometimes pathogenic. Our bodies have a symbiotic relationship with the “good guys” — some microbes break down the foods we eat, and others metabolize the drugs we take. But we are just beginning to uncover the interactions between metabolites released by microbes and the host’s response. How does the host deal with a small compound released by a bacterium? What methods are currently available to catalogue and study these interactions? And how can this interplay be used to combat gastrointestinal diseases and others affected by these metabolites? The recent minireview series “Host-microbiome metabolic interplay” in the Journal of Biological Chemistry seeks to address these questions, exploring the advanced genomics methods used today and examples of such interactions to illustrate their connection to human disease.

In compiling the work for this series, editor Ruma Banerjee of the University of Michigan notes, she chose scientists who explored the “sub-areas” of microbial metabolomics in their research. The finished result gives the reader six articles that cover several connected aspects of this field. The first two highlight three “-omics” methods used and several natural products uncovered with these tools, while the last four get into specific examples of microbial metabolites and the host’s response.
Emily Balskus and colleagues at Harvard University examine the workflow of advanced genomics techniques used to find natural products. Metagenomics, ecology-based approaches and in vitro biochemistry techniques were utilized in these case studies. The natural product and antibiotic lugdunin was discovered using an ecology-based approach in which a particular bacterial strain initially was found to inhibit the well-known pathogen Staphylococcus aureus. The cluster of genes correlating with this observation was identified and the specific metabolite isolated.
In the second article, the Eugene Chang laboratory of the University of Chicago delves into “-omics” methods. They explain how metagenomics, metatranscriptomics and metabolomics alone or in combination ultimately can aid in discovering therapies. Metagenomics, a mainstay of microbiome-host studies, uses shotgun sequencing to find novel material. Metatranscriptomics gives gene expression data, and metabolomics looks at host metabolic pathway changes. Inflammatory bowel disease is discussed as an example; in particular, metagenomic analysis revealed lowered microbial diversity in patients, which may contribute to the disease. In addition, metabolomics studies revealed differing levels of key microbial metabolites in IBD patients compared to controls.
The remaining articles describe specific examples of how metabolites from microbes interact with the host and the pros and cons of such close relationships. J. Mark Brown and Stanley Hazen of the Cleveland Clinic highlight the importance of targeting metabolites released from microbes instead of the microbes themselves. They chronicle the identification and inhibition of trimethylamine N-oxide (or TMAO, a biomarker for cardiovascular diseases) as a paradigm for this approach.
The fourth article addresses drug metabolism. Our bodies add sugars called glucuronides to drugs, which aids in the drugs’ elimination. Microbes, however, remove these sugars to use as an energy source. This puts drugs back into circulation and renders them toxic. Samuel Pellock and Matthew Redinbo of the University of North Carolina examine the types of molecules that glucuronides are added to and how microbes disturb this process.
Andreas Bäumler and colleagues at the University of California introduce the concept of colonization resistance and its disruption. Administration of antibiotics initially prevents bacteria from colonizing; yet after stopping the antibiotics or through antibiotic-resistant microbes, the colonization resistance is disrupted. The fifth article discusses what part of the microbial environment leads to a bacterium’s rapid flourishing. The authors introduce the nutrient niche hypothesis, which posits that a microbe can outgrow its competitors when the one nutrient it needs most is abundant. Enterobacteriaceae, present in hospitals nationwide, is one such example described here, and oxygen is that nutrient.
The last article describes an additional layer of control: epigenetics. Kimberly Krautkramer, Federico Rey and John Denu of the University of Wisconsin describe the modifications that occur on the histones that organize DNA in the nucleus and how microbial metabolites potentially could influence gene expression levels.
Banerjee says this field is an “interesting frontier for understanding metabolism as it is regulated by the microbiome.” The study of microbial metabolomics is in its early stages, and these articles are “beginning to scratch beyond the surface.” Banerjee hopes this will spark more in-depth studies using classic biochemistry coupled with advanced genomic methods to catalogue the interactions between microbial metabolites and host responses, as much of this interplay stems directly from the foods we eat and drugs and supplements we take.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
Using 'nature’s mistakes' as a window into Lafora disease
After years of heartbreak, Lafora disease families are fueling glycogen storage research breakthroughs, helping develop therapies that may treat not only Lafora but other related neurological disorders.

Cracking cancer’s code through functional connections
A machine learning–derived protein cofunction network is transforming how scientists understand and uncover relationships between proteins in cancer.

Gaze into the proteomics crystal ball
The 15th International Symposium on Proteomics in the Life Sciences symposium will be held August 17–21 in Cambridge, Massachusetts.
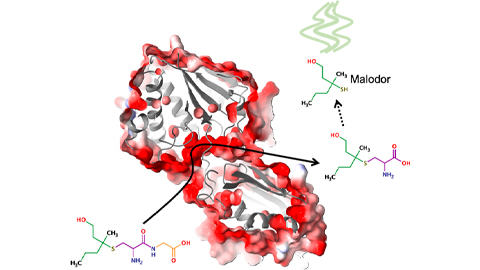
Bacterial enzyme catalyzes body odor compound formation
Researchers identify a skin-resident Staphylococcus hominis dipeptidase involved in creating sulfur-containing secretions. Read more about this recent Journal of Biological Chemistry paper.

Neurobiology of stress and substance use
MOSAIC scholar and proud Latino, Bryan Cruz of Scripps Research Institute studies the neurochemical origins of PTSD-related alcohol use using a multidisciplinary approach.
Pesticide disrupts neuronal potentiation
New research reveals how deltamethrin may disrupt brain development by altering the protein cargo of brain-derived extracellular vesicles. Read more about this recent Molecular & Cellular Proteomics article.

