Sharing the whole HeLa genome
In March 2013, a group of researchers at the European Molecular Biology Laboratory sequenced the genome of HeLa cells. With the last decades’ advances in sequencing techniques, the sequencing was done easily. It was also done with good intentions.
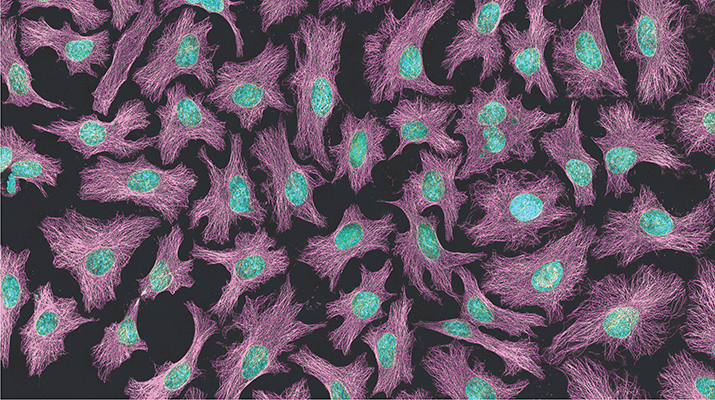
The cancer cells, which were first taken from a lump removed from Henrietta Lacks’ cervix months before her death from cervical cancer in 1951, are the most widely used cell line in the world. The cells are hardy and have helped develop many antitumor and viral treatments, including the polio vaccine. However, the genomic data published in 2013, which can be used to glean sensitive medical information about Lacks’ descendants, was shared without their knowledge.
“It’s like, ‘Here we go again, being involved in research without our permission or our consent,’” says David Lacks Jr. He is a grandson of Henrietta Lacks, who was a black tobacco farmer and a mother of five. When Henrietta Lacks went to seek medical attention at Johns Hopkins Hospital for a small mass in her cervix in 1951, the gynecologist on duty, Howard Jones, took a biopsy of the tumor cells. After a diagnosis, the cells made their way to George Gey, the head of tissue culture research at Johns Hopkins, by way of a mutual colleague.
Henrietta Lacks wasn’t asked for permission for her cells to be shared in this manner, although taking samples from patients without permission was a standard practice at the time. While her cells, which divided indefinitely at an unprecedented rate, went on to revolutionize medical research, the Lacks family was kept in the dark until researchers came looking to draw blood samples from family members in the 1970s. The HeLa cells generated billions of dollars of profit for biomedical industries, while the Lacks family was unable to afford medical care and health insurance.
These injustices were brought to the world’s attention with Rebecca Skloot’s bestselling 2010 book, “The Immortal Life of Henrietta Lacks.” Before publishing the book, Skloot established the Henrietta Lacks Foundation, which now has awarded more than 50 grants for education-related, health-care and pre-approved emergency expenses to a number of members of Lacks’ immediate family.
When the genome was put up on the European Nucleotide Archive in early 2013, “there weren’t any policies that said the data couldn’t be made available,” says Dina Paltoo at the National Institutes of Health. Paltoo is the director of the scientific data sharing policy division at the NIH’s office of science policy. “This is pretty much the standard practice in the genomics community, and a lot of journals require that data has been shared before they’ll publish the findings.” A study about the genome and epigenome of the HeLa cells by researchers at the University of Washington was also about to be published in Nature.
After the genomic information was put up in a public database by the German researchers at EMBL, Skloot published an op-ed in the New York Times that garnered a significant amount of attention. NIH Director Francis S. Collins met up with the Lacks family to discuss their options.
“We could leave it out there as is, for the whole world to see, but the issue with that is when you sequence Henrietta Lacks’ genome, you also include family traits of our genome as well,” says Lacks. “We don’t know what would be known 20 years from now with that sequence just being out there for anybody to use and how that would have an effect on us.”
Reaching a consensus
The family came to the conclusion that the best way to handle the HeLa genomic sequence would be to have researchers apply to access it. “We didn’t want it to be cut off, because the family is unanimously proud of what the cells have helped accomplish,” says Lacks.
Collins and Kathy Hudson, who then was the NIH’s deputy director for science, outreach and policy, put together a working group consisting of bioethicists, geneticists, clinicians and members of the Lacks family. According to the terms of the agreement in August 2013 that the family reached with the NIH, any researchers’ plans to use the data had to meet certain criteria: The data should be used only for biomedical research purposes, the requesters must disclose any commercial plans that they would have for the data, and the requesters would agree to acknowledge the family and the contributions of the cells in any publications and presentations. The study from the University of Washington group, which had been put on hold, appeared in an issue of Nature that ran that month with a discussion of the agreement by Hudson and Collins.
The HeLa Genome Data Access Working Group and includes Lacks and Veronica Spencer, a great-grandchild of Henrietta Lacks. The group evaluates requests to access this data and then sends its findings to the advisory committee to the NIH director. That committee then makes a recommendation to Collins, who makes a final decision.
“The NIH director has also reached out to journals and has encouraged them to make sure that investigators that are pursuing publication are abiding by the HeLa genome data use agreement and are also acknowledging the agreement and the family appropriately,” says Paltoo.
 David Lacks Jr. (right) and his cousin Jeri Lacks–Whye often speak publically about the Lacks family’s experiences with the HeLa cell line.PHOTO PROVDED BY JERI LACKS-WHYE
David Lacks Jr. (right) and his cousin Jeri Lacks–Whye often speak publically about the Lacks family’s experiences with the HeLa cell line.PHOTO PROVDED BY JERI LACKS-WHYE
Fruits of the database
The NIH’s database of genotypes and phenotypes, or dbGaP, currently contains five data sets related to the sequenced HeLa genome. So far, Collins has approved 47 requests from researchers from 20 different countries. The only rejected request was for a group that didn’t want to share its findings. The two papers that caused the uproar were published after they were approved by the group.
One of those approved investigators is Andrew Adey at Oregon Health & Science University. As a graduate student, Adey was the first author on the University of Washington genome paper led by Jay Shendure.
Early in his career, Adey helped investigate what gives the HeLa cells the ability to divide in such an aggressive manner. The capability arose from the integration of DNA from the human papilloma virus into the genome of a cell in Henrietta Lacks that led to her cervical carcinoma.
“The viral foreign DNA integration that occurred in the HeLa genome happens in some subset of cervical carcinomas, but in this case it happened in a very unfortunate way,” says Adey. “It happened to integrate in a location that activates a cancer gene, so it was really a perfect storm of events that happened in the cell that resulted in this extremely aggressive form of cancer and, ultimately, immortalization of the cell.”
The E6 and E7 viral oncogenes were present on the inserted viral DNA that inhibit tumor suppressors, such as the well-known p53. The virus also inserted 30 copies of a regulatory enhancer near a proto-oncogene, MYC, which can cause unregulated cell division when hijacked. This interaction contributed to a much more aggressive form of cancer.
Adey and colleagues recently characterized the stability and heterogeneity of HeLa cells using a technique called combinatorial indexing. The technique allows them to perform single-cell whole-genome sequencing at a higher throughput than was previously possible by barcoding individual cells.
The researchers first applied the technique to cancer cells from an advanced adenocarcinoma and were able to identify subpopulations within the tumor. In future uses, “we’ll be able to sample very low abundance subpopulations,” says Adey. “We might be able to then infer and detect some aspects that could be targetable in a different way than the rest of the tumor.”
In addition to all of the lifesaving medicines developed with HeLa cells, researchers trying to develop new medical technologies can use the HeLa genome as a powerful calibration tool.
“We’re developing new technologies and tools to look at cancer as well as other aspects or other diseases,” says Adey. “When we develop these tools, we want to test them out on something where we know the answer, so that’s what we use HeLa for. We know exactly what it’s going to look like.”
Controlled access to the HeLa genomic data has also resulted in the development of a new analytical method by Shendure’s group. The method involves chromosome-scale scaffoldings to assemble highly contiguous genomes from short reads. The reassembly is made possible by an algorithm that clusters fragments of the genome based on chromatin interaction data sets, which are useful for assigning, ordering and orienting the genomic sequences to chromosomes. The researchers first described the method, for which Shendure has also filed a patent, in a paper in the journal Nature Biotechnology in November 2013. In the paper, the researchers used the HeLa genome as one way to test the method to find interchromosomal rearrangements in cancer genomes.
Additionally, new insights into the effect of the genome’s spatial organization on transcription, which has significant implications for aberrations that occur in diseases, have been made by Yijuan Ruan’s group at The Jackson Laboratory Cancer Center in Bar Harbor, Maine.
While researchers use the HeLa cells to better understand countless aspects of cell biology, Lacks and Jeri Lacks–Whye, another one of Henrietta Lacks’ grandchildren, have traveled to speak to audiences of up to 4,000 about their family and the broader issues raised in Skloot’s book.
“Even though we speak a lot on the book, we’re also starting to speak more on the issues that are encompassed in the book, like health, prosperity and precision medicine,” says Lacks.
“Everybody is going to be sick at some point in time or affected by somebody who’s sick,” he adds. “We want to help scientists find cures.”
The top image, which is the same image shown on this month's cover, is a multiphoton fluorescence image of HeLa cells. Microtubules are in magenta; DNA is in cyan. Image is courtesy of Tom Derrinck at the National Center for Microscopy and Imaging Research.

Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
Using 'nature’s mistakes' as a window into Lafora disease
After years of heartbreak, Lafora disease families are fueling glycogen storage research breakthroughs, helping develop therapies that may treat not only Lafora but other related neurological disorders.

Cracking cancer’s code through functional connections
A machine learning–derived protein cofunction network is transforming how scientists understand and uncover relationships between proteins in cancer.

Gaze into the proteomics crystal ball
The 15th International Symposium on Proteomics in the Life Sciences symposium will be held August 17–21 in Cambridge, Massachusetts.
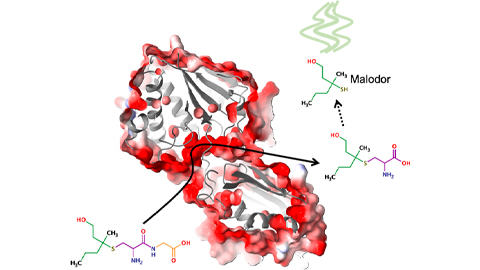
Bacterial enzyme catalyzes body odor compound formation
Researchers identify a skin-resident Staphylococcus hominis dipeptidase involved in creating sulfur-containing secretions. Read more about this recent Journal of Biological Chemistry paper.

Neurobiology of stress and substance use
MOSAIC scholar and proud Latino, Bryan Cruz of Scripps Research Institute studies the neurochemical origins of PTSD-related alcohol use using a multidisciplinary approach.
Pesticide disrupts neuronal potentiation
New research reveals how deltamethrin may disrupt brain development by altering the protein cargo of brain-derived extracellular vesicles. Read more about this recent Molecular & Cellular Proteomics article.

