JLR: Arteries and bacterial lipids don’t mix
Perhaps you have heard that brushing your teeth can be good for your heart. Bacteria of the Bacteroidetes phylum may be the main culprits behind the association between periodontal disease and atherosclerosis. In a recent paper in the Journal of Lipid Research, Frank Nichols and colleagues from the University of Connecticut reported on their investigation into how intestinal and oral Bacteriodetes may contribute to the development of atherosclerosis.
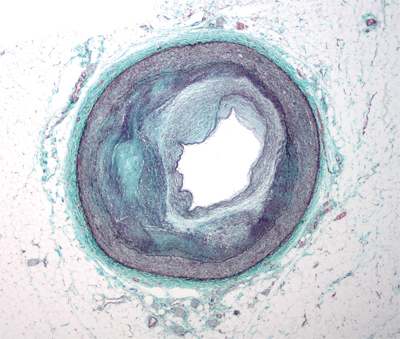 This low magnification micrograph shows the distal right coronary artery with complex atherosclerosis and luminal narrowing. The artery’s three layers can be seen as well as features of atherosclerosis. The section of the artery shown has only 25 to 35 percent of the cross-sectional area it once had. courtesy of WIKIMEDIA COMMONS/NEPHRON
This low magnification micrograph shows the distal right coronary artery with complex atherosclerosis and luminal narrowing. The artery’s three layers can be seen as well as features of atherosclerosis. The section of the artery shown has only 25 to 35 percent of the cross-sectional area it once had. courtesy of WIKIMEDIA COMMONS/NEPHRON
Atherosclerosis is a disease in which plaques build up in arteries. The plaques contain fat, cholesterol, calcium and other substances found in the blood. Over time, these plaques cause arteries to narrow and harden, and this leads to an increased risk of cardiovascular disease. Many factors can contribute to the development of atherosclerosis, including genetic predisposition, lipid metabolism and inflammation.
For their work, Nichols and his team first focused on a specific oral Bacteriodetes organism, Porphyromonas gingivalis. This organism has been associated with an increased risk of developing atherosclerosis in rodents and periodontal disease in humans (1,2). In addition, Nichols’ group has shown that both oral and intestinal Bacteroidetes produce specific serine dipeptide lipids. The group also has shown that these bacterial lipids activate macrophages and immune cells that promote inflammation in tissues.
Two serine dipeptide lipids are produced by P. gingivalis, Lipid 654 and Lipid 430. A chemical bond in Lipid 654 is cleaved to yield Lipid 430.
For this study, the group relied on high-performance liquid chromatography, or HPLC, and tandem mass spectrometry to isolate and quantify Lipid 430 to Lipid 654 ratios. HPLC technology can resolve component lipids of a complex lipid mixture using a column containing particles of silica gel. These particles can interact with, in this case, lipid components. As the lipid components elute from the HPLC column, the eluting lipids can be introduced into a mass spectrometer to identify specific metabolites and quantify their levels in a given sample. Using this technology, the researchers saw not only that Lipid 654 was present in human carotid arteries but also that the Lipid 430 to Lipid 654 ratio increased by at least tenfold in diseased arteries compared with artery samples from young subjects. This suggests that Lipid 654 hydrolysis is increased in diseased carotid arteries. “Many think that atherosclerosis is caused by eating fatty foods,” Nichols said, “but it is now apparent that other lipids produced by oral and intestinal bacteria accumulate in diseased arteries.”
In addition to finding an accumulation of Lipid 430 in diseased human carotid arteries, the group identified a mechanism for cleavage of Lipid 654. They tested a panel of common lipase enzymes, some of which are associated with atherosclerosis, for their ability to hydrolyze Lipid 654 to Lipid 430. They tested human, pig or honey bee enzymes. After incubation with the enzymes, the amounts of Lipid 654 and Lipid 430 were measured by mass spectrometry. They found that phospholipase A2, an enzyme associated with macrophage activation, was the only enzyme class in the panel that was able to cleave Lipid 654.
The research by Nichols’ lab suggests that patients with atherosclerotic disease have an elevated level of Lipid 430 resulting from Lipid 654 cleavage mediated by phospholipase A2. Furthermore, Lipid 430 potentially can promote inflammation at the site of plaques. “For the future,” Nichols said, “it will be important to identify the enzymes responsible for the synthesis of these lipids and target specific enzymes with pharmacological inhibitors and/or dietary modifications.”
REFERENCES
1. Gibson III, F.C., et al. Curr. Pharm. Des. 13, 3665-3675 (2007).
2. Armingohar, Z., et al. J. Oral Microbiol. 6, 23408 (2014).
Enjoy reading ASBMB Today?
Become a member to receive the print edition monthly and the digital edition weekly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
Iron could be key to treating a global parasitic disease
A study has found that leishmaniasis causes body-wide changes in iron balance, leading to red blood cell damage.

Environmental DNA is everywhere
The ability to extract trace bits of DNA from soil, water, and even air is revolutionizing science. Are there pitfalls?
Early COVID-19 research is riddled with poor methods and low-quality results
The pandemic worsened, but didn’t create, this problem for science.

From the journals: MCP
Three views of mass spec: analyzing secreted protein spectra, imaging mass spectrometry for clinical use and spectral libraries for MS data analysis. Read about these recent papers.

Understanding the fat science
Researchers at UCLA investigate lipid remodeling in the liver for energy generation.

No oxygen? No problem
By studying how electric fish survive in hypoxic streams for months at time, researchers may find new ways to target tumors.

